library(tidyverse)
theme_set(theme_light())
library(magrittr)
library(coretta2018itaegg)
library(brms)
library(rstan)
library(posterior)
library(tidybayes)
library(ggdist)
library(mgcv)
library(tidygam)
library(HDInterval)
library(truncdist)
library(ggdist)
my_seed <- 9899
cols <- viridisLite::viridis(5)
cols[5] <- "#d95f02"Analysis of intrinsic vowel duration in Northwestern Italian
1 Attach packages
2 Read data
The following code loads the data from the coretta2018itaegg package and creates new variables as transformations of the existing ones.
data("ita_egg")
ita_egg <- ita_egg |>
drop_na(f13, f23) |>
mutate(
duration = v1_duration,
vowel = as.factor(vowel),
c1_place = as.factor(c1_place),
c2_place = as.factor(c2_place),
duration_z = as.vector(scale(duration)),
duration_log = log(duration),
duration_logz = as.vector(scale(log(duration))),
speech_rate_log = log(speech_rate),
speech_rate_logz = as.vector(scale(log(speech_rate_log))),
f13_z = as.vector(scale(f13)),
f23_z = as.vector(scale(f23)),
speaker = as.factor(speaker),
word_ipa = str_replace_all(word, "ch|c", "k"),
previous_sound = str_sub(word_ipa, 1, 1),
next_sound = str_sub(word_ipa, 3, 3)
)3 Plotting
ita_egg |>
group_by(speaker) |>
mutate(
f13_speaker_z = as.vector(scale(f13)),
f23_speaker_z = as.vector(scale(f23))
) |>
ggplot(aes(f23_speaker_z, f13_speaker_z, colour = vowel)) +
geom_point(alpha = 0.2) +
stat_ellipse(type = "norm") +
scale_x_reverse(position = "top") + scale_y_reverse(position = "right") +
coord_fixed()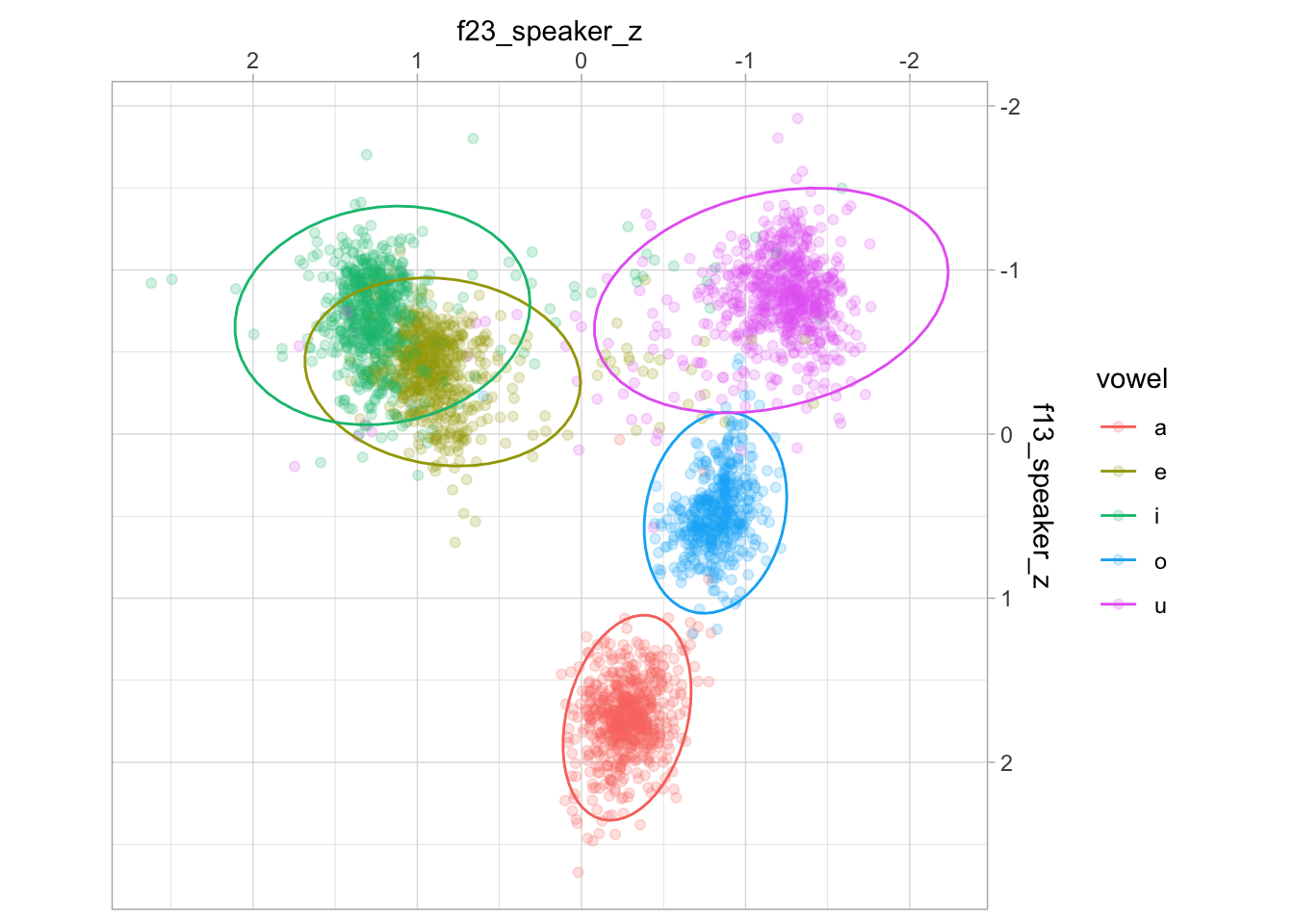
ita_egg |>
group_by(speaker) |>
mutate(
f13_speaker_z = as.vector(scale(f13)),
f23_speaker_z = as.vector(scale(f23))
) |>
ggplot(aes(f23_speaker_z, f13_speaker_z)) +
geom_point(aes(colour = duration), alpha = 0.8) +
stat_ellipse(aes(group = vowel), type = "norm") +
scale_x_reverse(position = "top") + scale_y_reverse(position = "right") +
coord_fixed()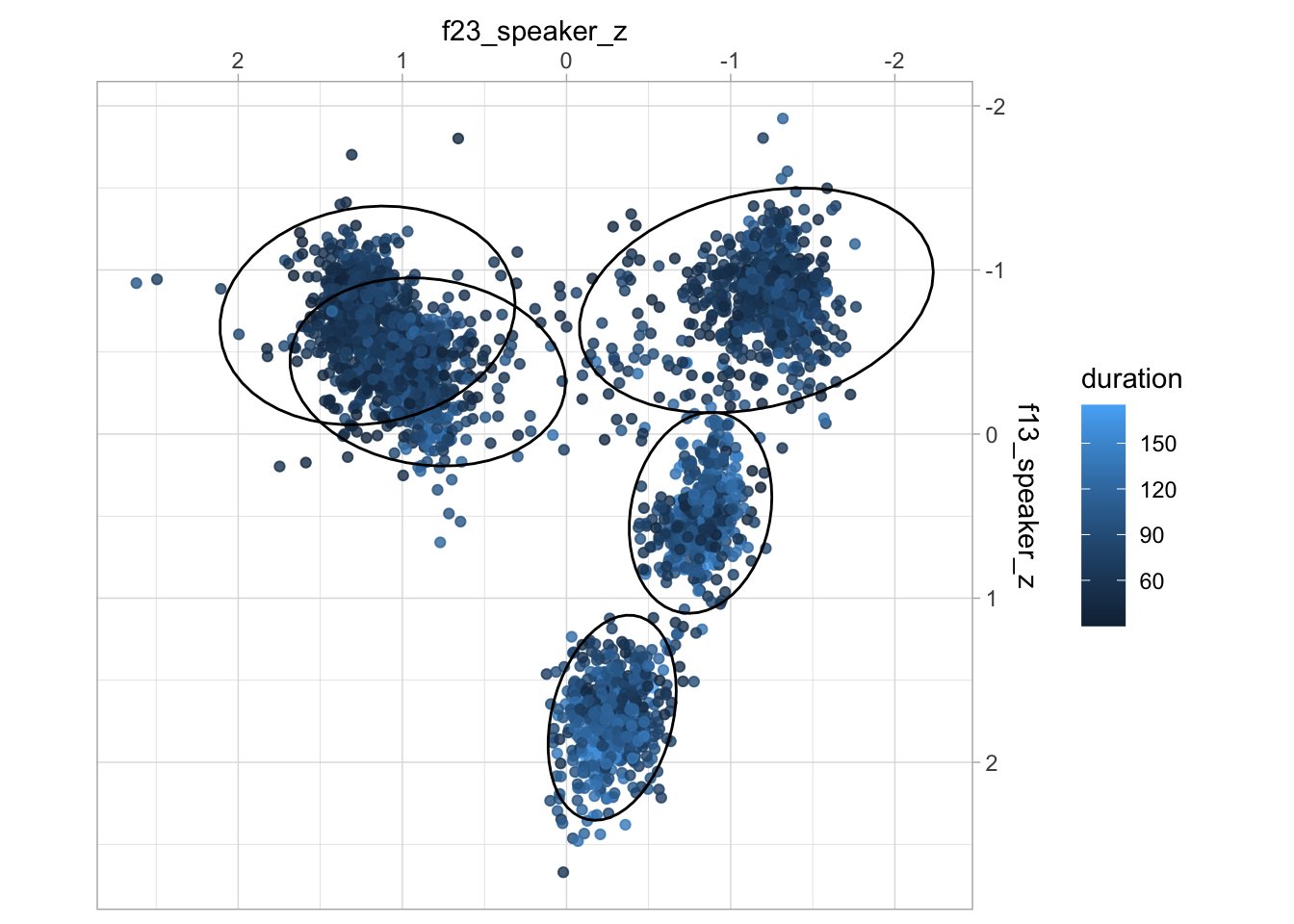
ita_egg |>
ggplot(aes(f13_z, duration_z)) +
geom_density_2d_filled()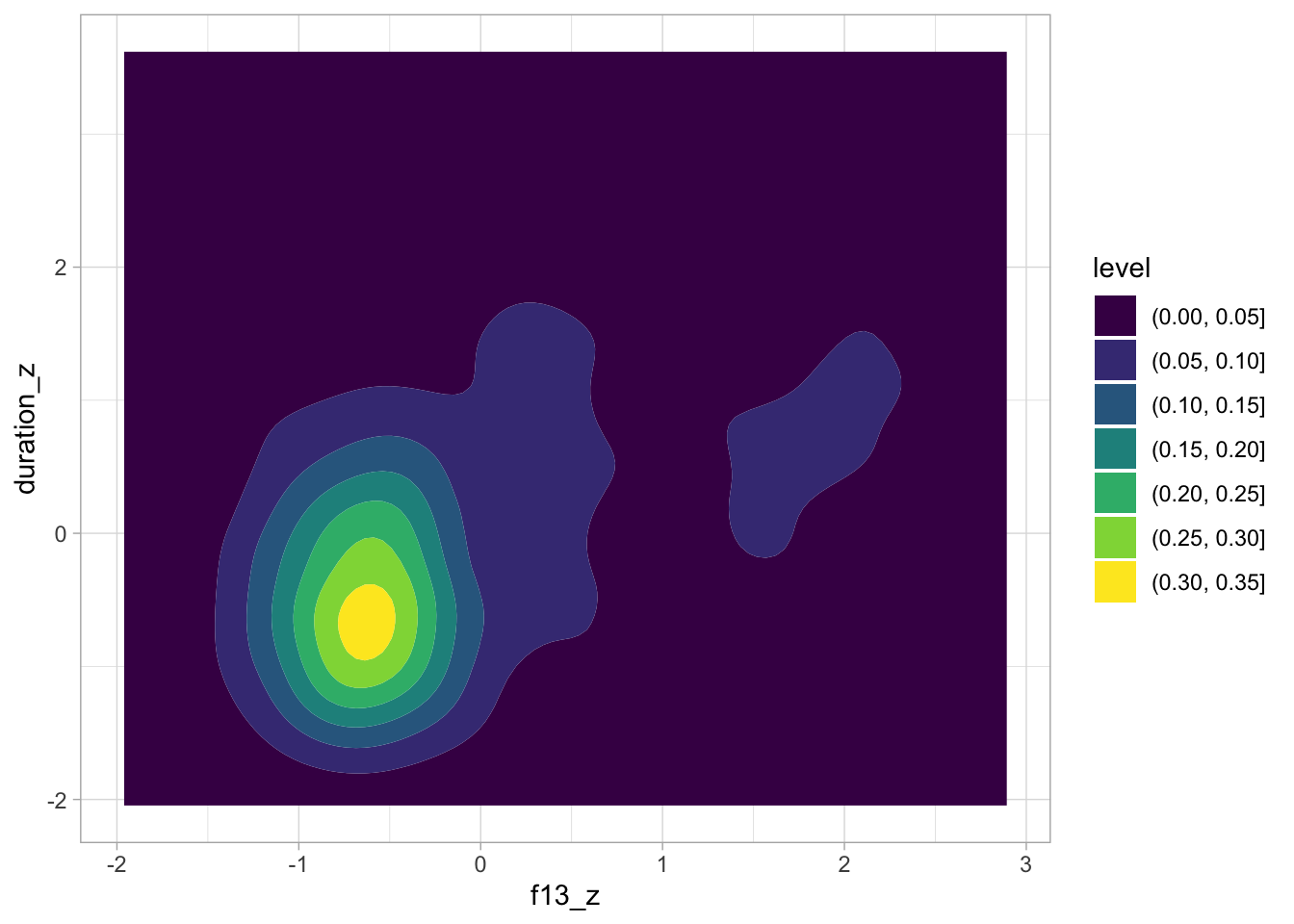
ita_egg |>
ggplot(aes(vowel, duration)) +
geom_jitter(width = 0.2, alpha = 0.2) +
facet_grid(rows = vars(next_sound), cols = vars(previous_sound))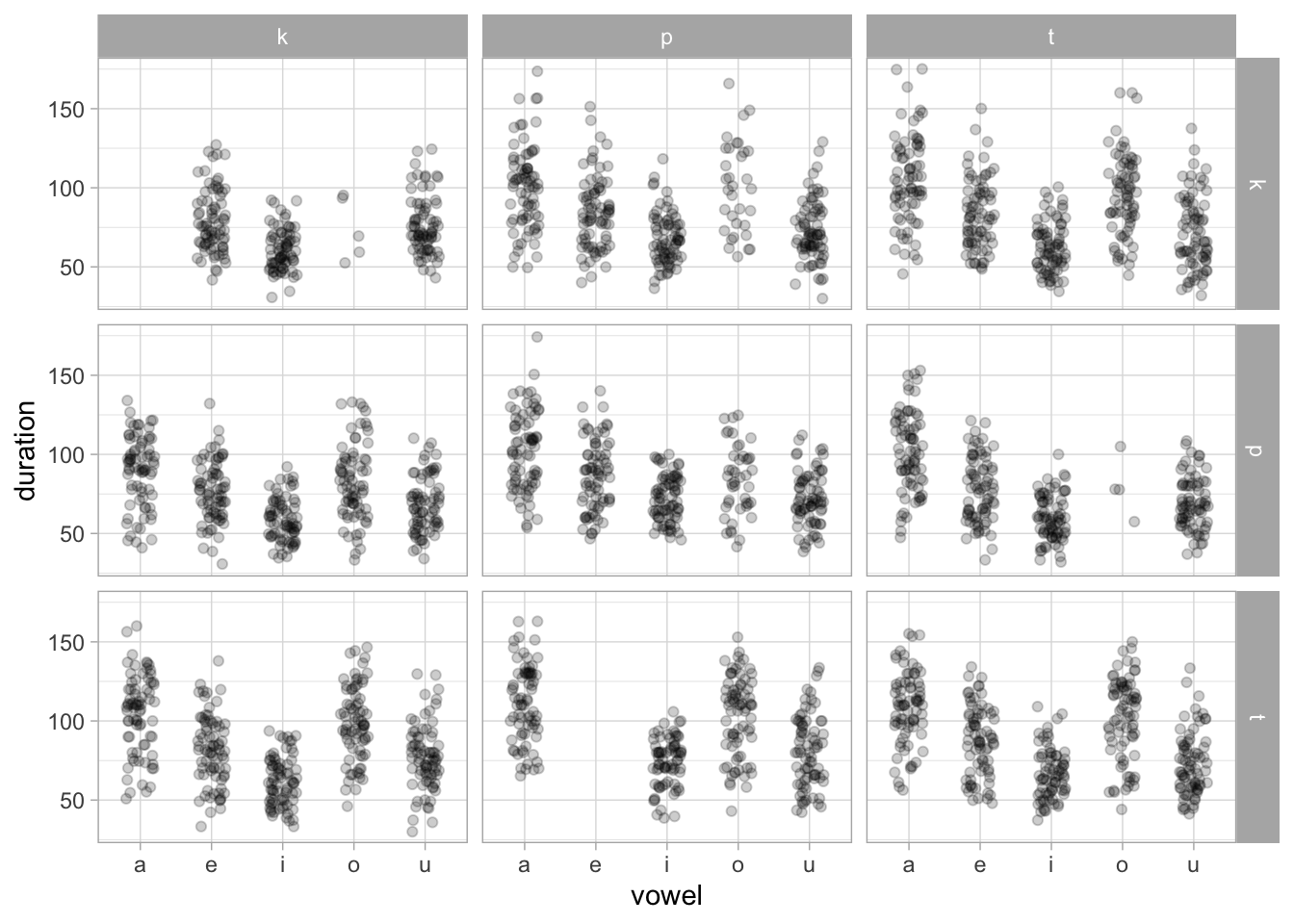
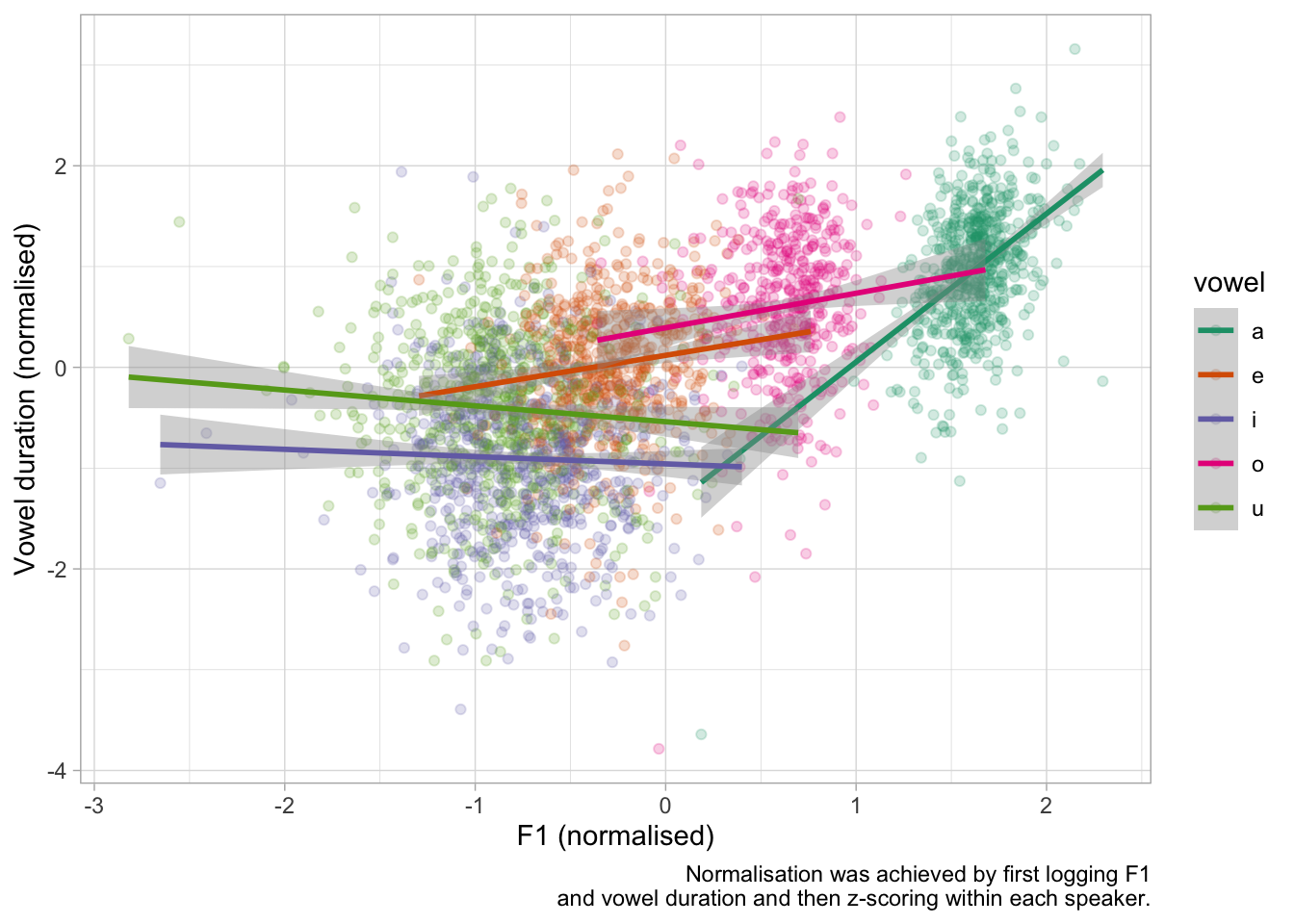
ita_egg |>
group_by(speaker) |>
mutate(
f13_speaker_z = as.vector(scale(log(f13))),
duration_speaker_z = as.vector(scale(log(duration)))
) |>
ggplot(aes(f13_speaker_z, duration_speaker_z, colour = vowel)) +
geom_point(alpha = 0.2) +
geom_smooth(method = "lm", formula = y ~ x) +
scale_color_brewer(type = "qual", palette = "Dark2") +
labs(
x = "F1 (normalised)",
y = "Vowel duration (normalised)",
caption = "Normalisation was achieved by first logging F1\nand vowel duration and then z-scoring within each speaker."
)
ggsave("img/dur-f1.png", width = 7, height = 5)4 Linear modelling
4.1 Prior predictive checks
The outcome duration_logz and predictor f13_z are z-scored and the intercept has been suppressed so that the indexing method is used for the vowel predictor instead of contrasts.
The following code sets up the model formula and gets the default priors. This is the mathematical representation of the model.
$$ \[\begin{align*} y_i &\sim \mathcal{N}(\mu_i, \sigma) \\ \mu_i &= a_i + b_i \cdot \text{F1}_i + c_i \cdot sr_i \\ a_i &= \alpha_{\text{vowel}[i], \,\text{c1}[i], \,\text{c2}[i]} + u^{(a)}_{\text{speaker}[i], \,\text{vowel}[i], \,\text{c1}[i], \,\text{c2}[i]} \\ b_i &= \beta_{\text{vowel}[i], \,\text{c1}[i], \,\text{c2}[i]} + u^{(b)}_{\text{speaker}[i], \,\text{vowel}[i], \,\text{c1}[i], \,\text{c2}[i]} \\ c_i &= \gamma + u^{(c)}_{\text{speaker}[i]} \\ \begin{bmatrix} u^{(a)}_{\text{speaker}[i], \cdot} \\ u^{(b)}_{\text{speaker}[i], \cdot} \end{bmatrix} &\sim \mathcal{N}\!\left( \mathbf{0}, \, \Sigma \right), \quad u^{(c)}_{\text{speaker}[i]} \sim \mathcal{N}(0, \tau_c^2) \end{align*}\]
$$
The \(a_i\) and \(b_i\) coefficients correspond to the by-vowel intercept and the F1 effect.
bm_1_f <- bf(
duration_logz ~ a + b*f13_z + c*speech_rate_logz,
a ~ 0 + vowel:c1_place:c2_place + (0 + vowel:c1_place:c2_place | speaker),
b ~ 0 + vowel:c1_place:c2_place + (0 + vowel:c1_place:c2_place | speaker),
c ~ (1 | speaker),
nl = TRUE
)
bm_1_get <- get_prior(
bm_1_f,
data = ita_egg,
family = gaussian,
) |> as_tibble()
bm_1_get# A tibble: 194 × 10
prior class coef group resp dpar nlpar lb ub source
<chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
1 "lkj(1)" cor "" "" "" "" "" "" "" defau…
2 "" cor "" "spe… "" "" "" "" "" defau…
3 "student_t(3, 0, 2.5)" sigma "" "" "" "" "" "0" "" defau…
4 "" b "" "" "" "" "a" "" "" defau…
5 "" b "vow… "" "" "" "a" "" "" defau…
6 "" b "vow… "" "" "" "a" "" "" defau…
7 "" b "vow… "" "" "" "a" "" "" defau…
8 "" b "vow… "" "" "" "a" "" "" defau…
9 "" b "vow… "" "" "" "a" "" "" defau…
10 "" b "vow… "" "" "" "a" "" "" defau…
# ℹ 184 more rowsWe fit the prior predictive model with relatively weakly informative priors. The prior predictive model is the model fitted with sample_prior = "only" so that only the priors are sampled, without including the data. This enables us to check the joint predictions based on priors only.
priors <- c(
prior(normal(0, 1), class = b, nlpar = a),
prior(normal(0, 1), class = b, nlpar = b),
prior(normal(0, 0.1), class = b, nlpar = c),
prior(cauchy(0, 0.1), class = sigma),
prior(lkj(2), class = cor),
prior(cauchy(0, 0.1), class = sd, nlpar = a),
prior(cauchy(0, 0.1), class = sd, nlpar = b),
prior(cauchy(0, 0.1), class = sd, nlpar = c)
)
bm_1_priors <- brm(
bm_1_f,
family = gaussian,
data = ita_egg,
prior = priors,
cores = 4,
init = 0,
threads = threading(2),
backend = "cmdstanr",
sample_prior = "only",
file = "data/cache/bm_1_priors",
)The following plots show the prior predictive values of vowel duration, based exclusively on the priors. The prior predictive plots indicate that the priors cover a reasonable range of values without being too informative.
conditional_effects(bm_1_priors, "f13_z:vowel", spaghetti = TRUE, ndraws = 50)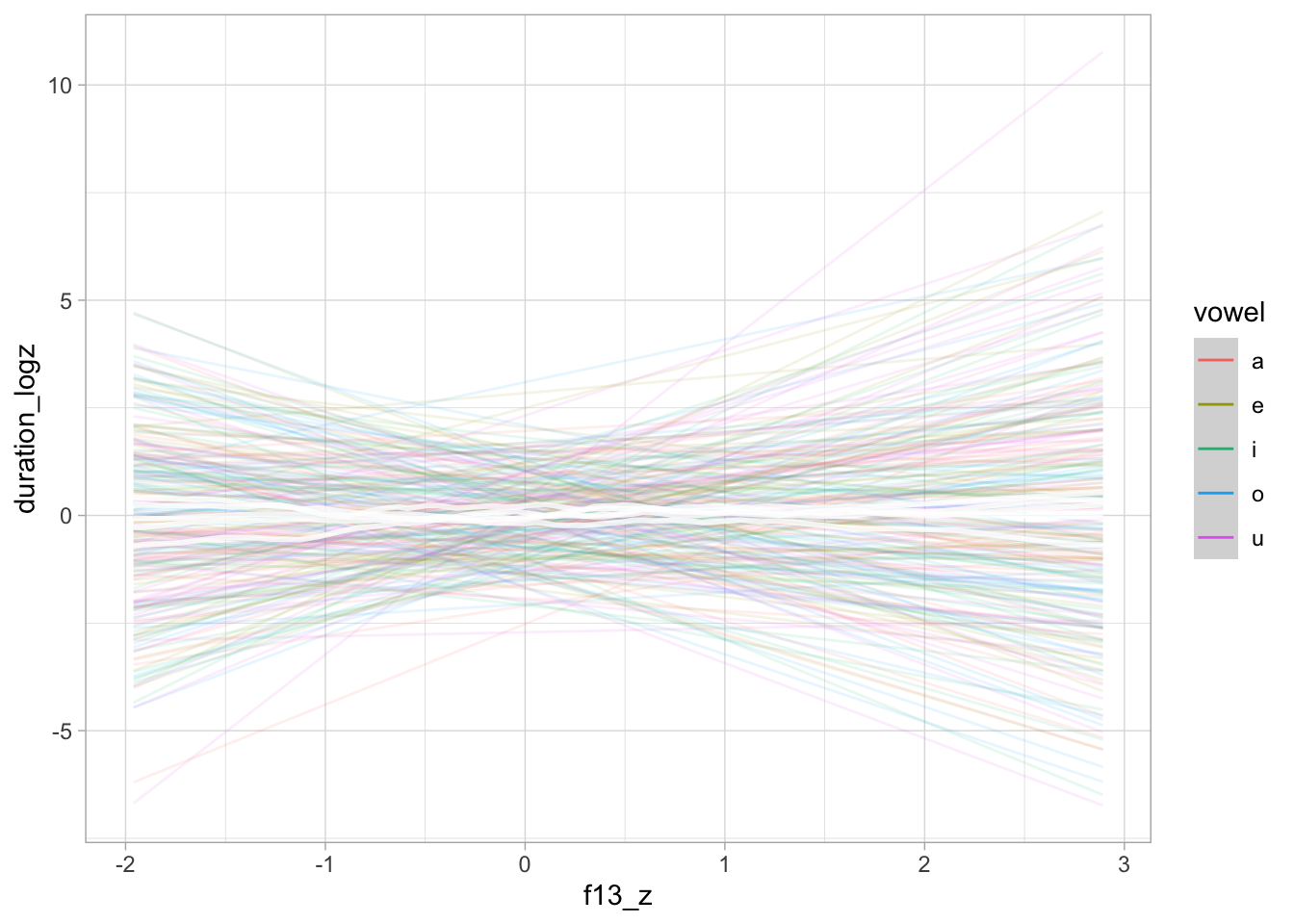
conditional_effects(bm_1_priors, "f13_z:c1_place", spaghetti = TRUE, ndraws = 50)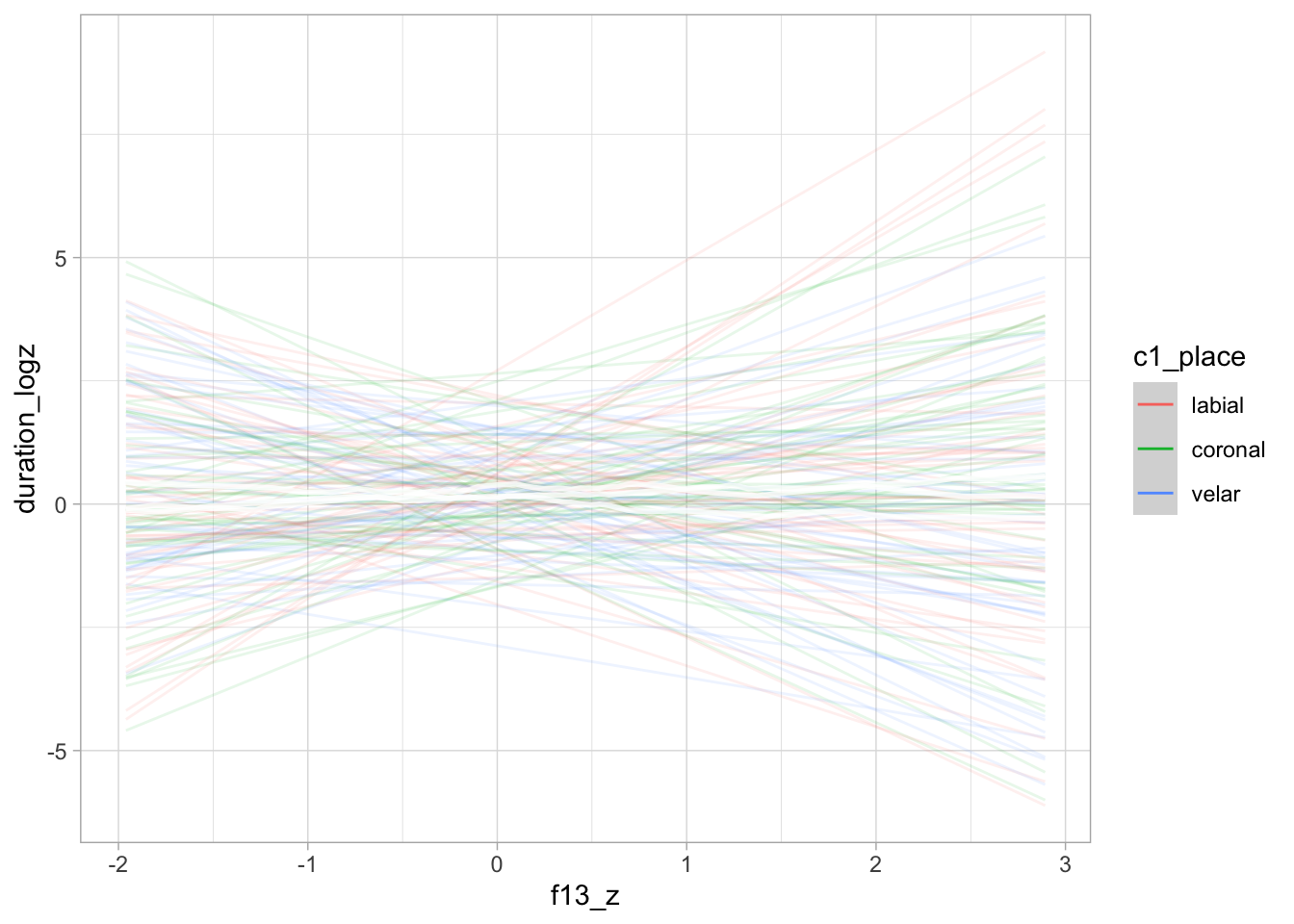
conditional_effects(bm_1_priors, "f13_z:c2_place", spaghetti = TRUE, ndraws = 50)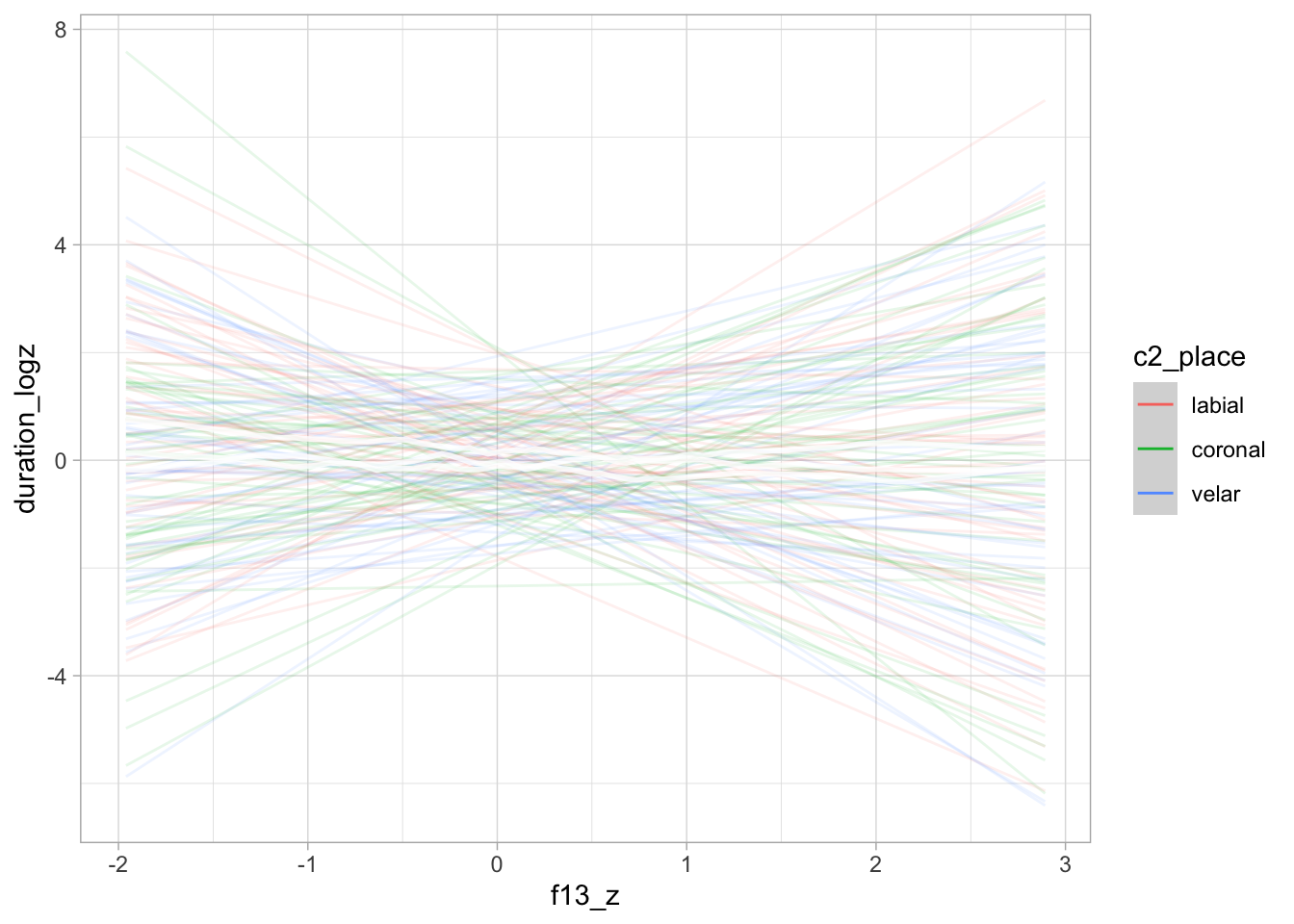
conditional_effects(bm_1_priors, "speech_rate_logz", spaghetti = TRUE, ndraws = 50)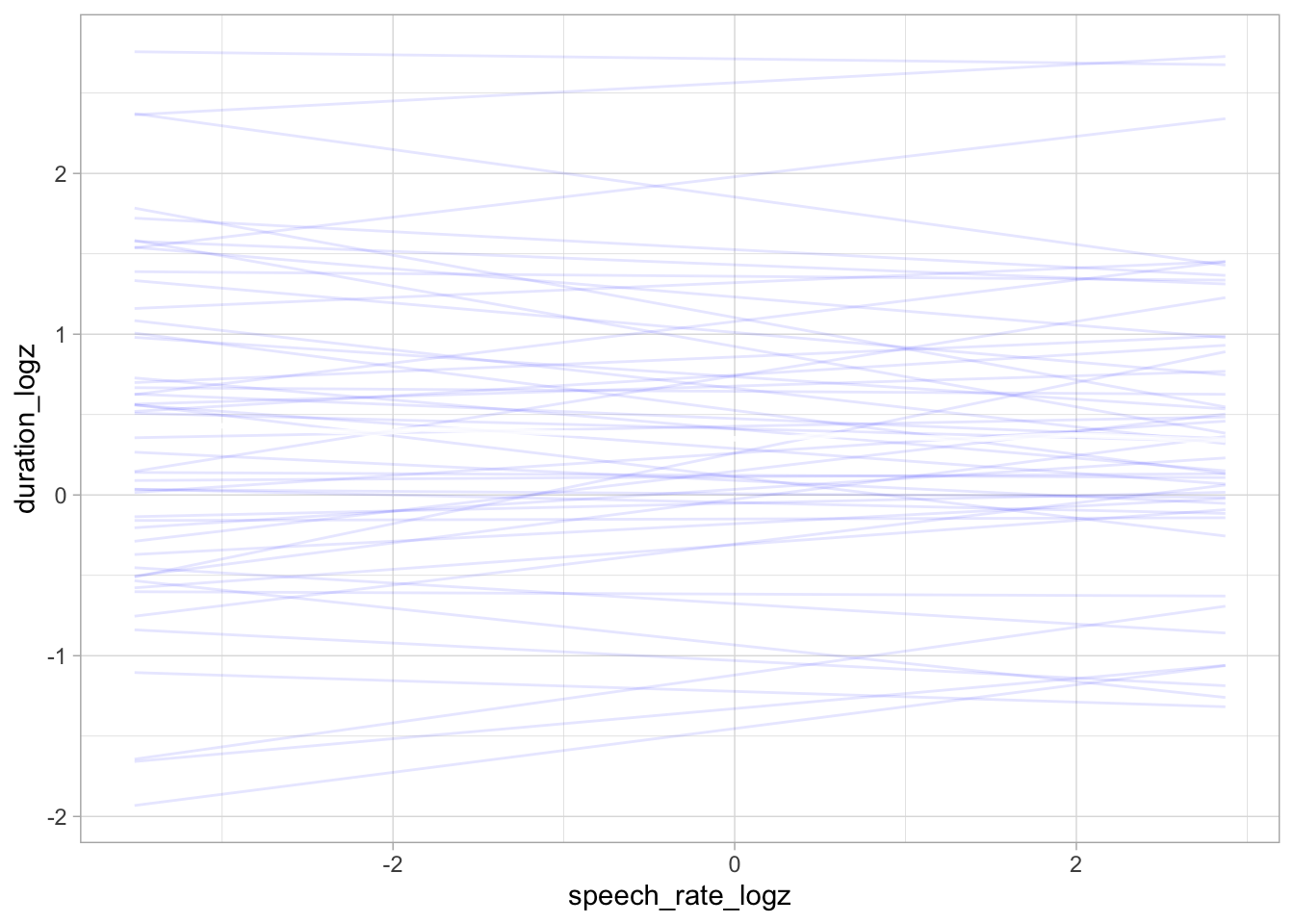
conditional_effects(bm_1_priors, "vowel")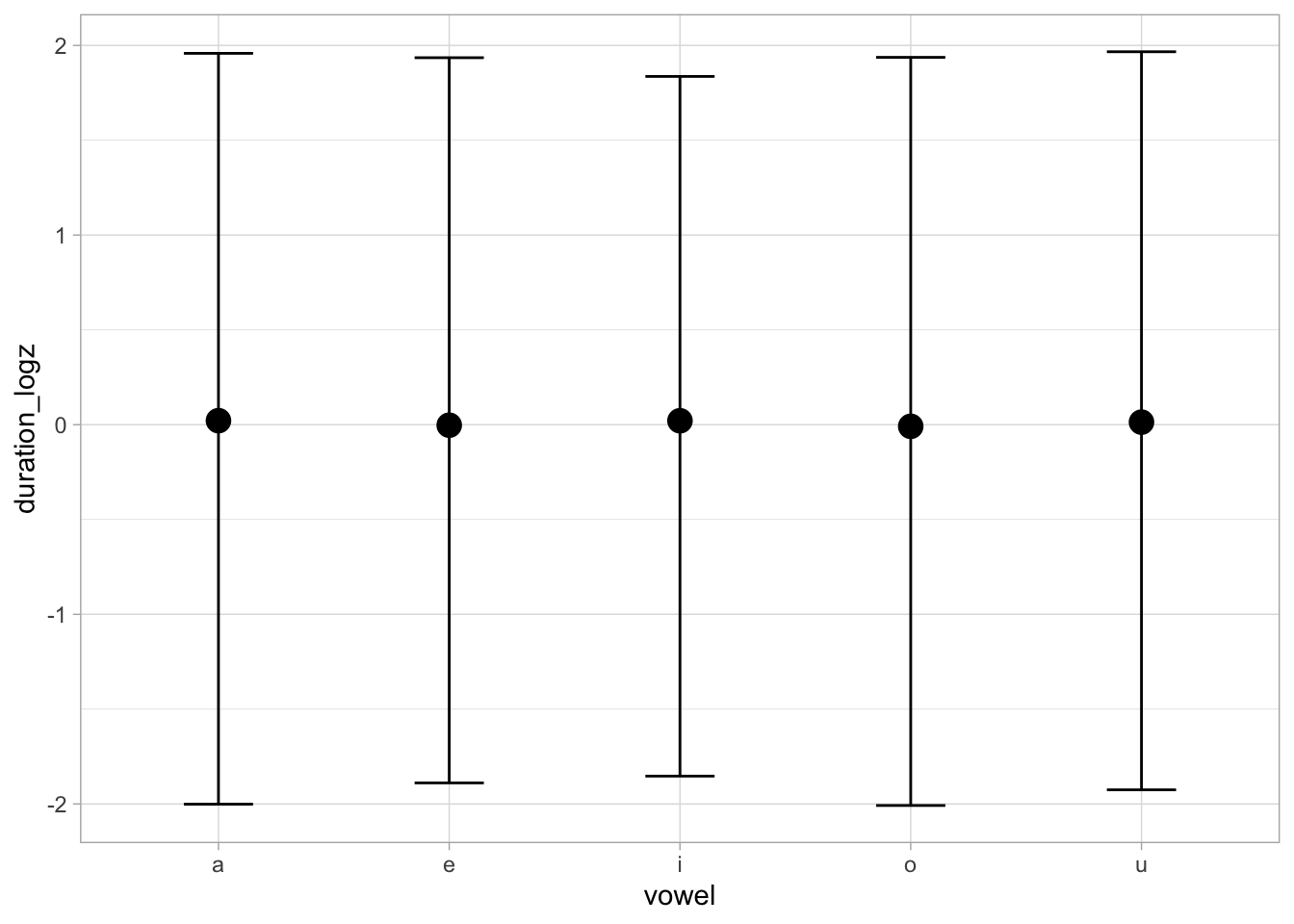
4.2 Model fit
We now fit the model with the specified priors to the data.
bm_1 <- brm(
bm_1_f,
family = gaussian,
data = ita_egg,
prior = priors,
init = 0,
cores = 4,
threads = threading(2),
backend = "cmdstanr",
file = "data/cache/bm_1",
)The following are the regression coefficients as estimated by the model.
fixef(bm_1, probs = c(0.05, 0.95)) |> round(2) Estimate Est.Error Q5 Q95
a_vowela:c1_placelabial:c2_placelabial -0.05 0.29 -0.55 0.42
a_vowele:c1_placelabial:c2_placelabial 0.38 0.13 0.17 0.59
a_voweli:c1_placelabial:c2_placelabial -0.31 0.13 -0.54 -0.09
a_vowelo:c1_placelabial:c2_placelabial 0.10 0.19 -0.21 0.41
a_vowelu:c1_placelabial:c2_placelabial -0.48 0.18 -0.77 -0.17
a_vowela:c1_placecoronal:c2_placelabial -0.12 0.26 -0.56 0.30
a_vowele:c1_placecoronal:c2_placelabial 0.10 0.15 -0.14 0.34
a_voweli:c1_placecoronal:c2_placelabial -0.89 0.18 -1.18 -0.61
a_vowelo:c1_placecoronal:c2_placelabial 0.19 0.29 -0.26 0.67
a_vowelu:c1_placecoronal:c2_placelabial -0.55 0.18 -0.84 -0.25
a_vowela:c1_placevelar:c2_placelabial -0.22 0.27 -0.69 0.22
a_vowele:c1_placevelar:c2_placelabial -0.03 0.13 -0.23 0.19
a_voweli:c1_placevelar:c2_placelabial -1.15 0.16 -1.41 -0.89
a_vowelo:c1_placevelar:c2_placelabial -0.07 0.14 -0.30 0.15
a_vowelu:c1_placevelar:c2_placelabial -0.46 0.23 -0.84 -0.08
a_vowela:c1_placelabial:c2_placecoronal 0.20 0.29 -0.29 0.68
a_vowele:c1_placelabial:c2_placecoronal 0.00 1.00 -1.65 1.65
a_voweli:c1_placelabial:c2_placecoronal -0.33 0.15 -0.57 -0.08
a_vowelo:c1_placelabial:c2_placecoronal 0.53 0.17 0.25 0.80
a_vowelu:c1_placelabial:c2_placecoronal -0.20 0.18 -0.50 0.10
a_vowela:c1_placecoronal:c2_placecoronal 0.38 0.30 -0.12 0.87
a_vowele:c1_placecoronal:c2_placecoronal 0.40 0.14 0.18 0.62
a_voweli:c1_placecoronal:c2_placecoronal -0.53 0.18 -0.83 -0.24
a_vowelo:c1_placecoronal:c2_placecoronal 0.38 0.16 0.13 0.64
a_vowelu:c1_placecoronal:c2_placecoronal -0.30 0.21 -0.64 0.04
a_vowela:c1_placevelar:c2_placecoronal 0.17 0.30 -0.33 0.65
a_vowele:c1_placevelar:c2_placecoronal 0.14 0.13 -0.07 0.36
a_voweli:c1_placevelar:c2_placecoronal -0.56 0.18 -0.85 -0.25
a_vowelo:c1_placevelar:c2_placecoronal 0.69 0.14 0.45 0.93
a_vowelu:c1_placevelar:c2_placecoronal 0.00 0.20 -0.33 0.32
a_vowela:c1_placelabial:c2_placevelar 0.34 0.33 -0.21 0.88
a_vowele:c1_placelabial:c2_placevelar 0.18 0.11 0.01 0.35
a_voweli:c1_placelabial:c2_placevelar -0.44 0.15 -0.69 -0.19
a_vowelo:c1_placelabial:c2_placevelar 0.55 0.20 0.21 0.87
a_vowelu:c1_placelabial:c2_placevelar 0.08 0.21 -0.25 0.43
a_vowela:c1_placecoronal:c2_placevelar 0.31 0.29 -0.18 0.78
a_vowele:c1_placecoronal:c2_placevelar 0.33 0.12 0.13 0.52
a_voweli:c1_placecoronal:c2_placevelar -0.74 0.16 -0.99 -0.48
a_vowelo:c1_placecoronal:c2_placevelar 0.43 0.15 0.18 0.67
a_vowelu:c1_placecoronal:c2_placevelar -0.58 0.20 -0.91 -0.25
a_vowela:c1_placevelar:c2_placevelar 0.01 1.02 -1.66 1.66
a_vowele:c1_placevelar:c2_placevelar 0.10 0.12 -0.10 0.30
a_voweli:c1_placevelar:c2_placevelar -0.88 0.14 -1.11 -0.65
a_vowelo:c1_placevelar:c2_placevelar -0.22 0.27 -0.64 0.21
a_vowelu:c1_placevelar:c2_placevelar -0.30 0.19 -0.60 0.02
b_vowela:c1_placelabial:c2_placelabial 0.49 0.17 0.22 0.77
b_vowele:c1_placelabial:c2_placelabial 0.46 0.25 0.05 0.89
b_voweli:c1_placelabial:c2_placelabial 0.05 0.17 -0.24 0.33
b_vowelo:c1_placelabial:c2_placelabial 0.17 0.36 -0.43 0.76
b_vowelu:c1_placelabial:c2_placelabial -0.16 0.21 -0.50 0.19
b_vowela:c1_placecoronal:c2_placelabial 0.50 0.15 0.27 0.75
b_vowele:c1_placecoronal:c2_placelabial 0.58 0.28 0.13 1.03
b_voweli:c1_placecoronal:c2_placelabial 0.03 0.22 -0.33 0.37
b_vowelo:c1_placecoronal:c2_placelabial 0.30 0.58 -0.66 1.20
b_vowelu:c1_placecoronal:c2_placelabial -0.11 0.19 -0.43 0.20
b_vowela:c1_placevelar:c2_placelabial 0.37 0.16 0.11 0.64
b_vowele:c1_placevelar:c2_placelabial 0.51 0.24 0.11 0.91
b_voweli:c1_placevelar:c2_placelabial -0.17 0.20 -0.50 0.16
b_vowelo:c1_placevelar:c2_placelabial 0.26 0.22 -0.09 0.62
b_vowelu:c1_placevelar:c2_placelabial 0.03 0.26 -0.39 0.47
b_vowela:c1_placelabial:c2_placecoronal 0.48 0.16 0.22 0.76
b_vowele:c1_placelabial:c2_placecoronal 0.00 0.99 -1.61 1.61
b_voweli:c1_placelabial:c2_placecoronal -0.12 0.20 -0.45 0.21
b_vowelo:c1_placelabial:c2_placecoronal 0.52 0.30 0.03 1.01
b_vowelu:c1_placelabial:c2_placecoronal -0.23 0.21 -0.58 0.11
b_vowela:c1_placecoronal:c2_placecoronal 0.35 0.17 0.07 0.63
b_vowele:c1_placecoronal:c2_placecoronal 0.42 0.25 0.01 0.83
b_voweli:c1_placecoronal:c2_placecoronal 0.08 0.21 -0.26 0.44
b_vowelo:c1_placecoronal:c2_placecoronal 0.73 0.24 0.33 1.12
b_vowelu:c1_placecoronal:c2_placecoronal 0.02 0.22 -0.33 0.37
b_vowela:c1_placevelar:c2_placecoronal 0.40 0.17 0.12 0.68
b_vowele:c1_placevelar:c2_placecoronal 0.40 0.24 0.01 0.80
b_voweli:c1_placevelar:c2_placecoronal 0.39 0.22 0.04 0.75
b_vowelo:c1_placevelar:c2_placecoronal -0.15 0.24 -0.55 0.25
b_vowelu:c1_placevelar:c2_placecoronal 0.23 0.21 -0.13 0.57
b_vowela:c1_placelabial:c2_placevelar 0.22 0.19 -0.09 0.55
b_vowele:c1_placelabial:c2_placevelar 0.22 0.21 -0.12 0.56
b_voweli:c1_placelabial:c2_placevelar 0.10 0.19 -0.21 0.41
b_vowelo:c1_placelabial:c2_placevelar -0.06 0.31 -0.57 0.46
b_vowelu:c1_placelabial:c2_placevelar 0.56 0.24 0.18 0.94
b_vowela:c1_placecoronal:c2_placevelar 0.30 0.16 0.04 0.57
b_vowele:c1_placecoronal:c2_placevelar 0.76 0.24 0.36 1.16
b_voweli:c1_placecoronal:c2_placevelar 0.13 0.21 -0.21 0.47
b_vowelo:c1_placecoronal:c2_placevelar 0.18 0.23 -0.21 0.55
b_vowelu:c1_placecoronal:c2_placevelar -0.21 0.22 -0.57 0.14
b_vowela:c1_placevelar:c2_placevelar -0.01 1.00 -1.66 1.65
b_vowele:c1_placevelar:c2_placevelar 0.57 0.23 0.19 0.96
b_voweli:c1_placevelar:c2_placevelar 0.04 0.19 -0.26 0.35
b_vowelo:c1_placevelar:c2_placevelar 1.12 0.63 0.02 2.10
b_vowelu:c1_placevelar:c2_placevelar -0.13 0.21 -0.47 0.22
c_Intercept -0.29 0.03 -0.34 -0.23We can now extract the MCMC draws of the \(\alpha\) and \(\beta\) coefficients for further processing. We convert the draws to a long format.
bm_1_draws <- as_draws_df(bm_1)
bm_1_draws_long <- bm_1_draws |>
select(
`b_a_vowela:c1_placelabial:c2_placelabial`:`b_b_vowelu:c1_placevelar:c2_placevelar`,
.chain, .iteration, .draw
) |>
pivot_longer(
`b_a_vowela:c1_placelabial:c2_placelabial`:`b_b_vowelu:c1_placevelar:c2_placevelar`
) |>
separate(name, c("vowel", "c1_place", "c2_place"), sep = ":") |>
separate(vowel, c("par", "coef", "vowel"))The following code summarises the draws for each coefficients, by vowel but averaged across the consonant place of articulation. a is the by-vowel intercept, while b is the effect of F1.
bm_1_draws_long_avg <- bm_1_draws_long |>
group_by(.chain, .iteration, .draw, coef, vowel) |>
summarise(epred = mean(value), .groups = "drop") |>
mutate(
vowel = str_remove(vowel, "vowel")
)
bm_1_draws_long_avg |>
group_by(coef, vowel) |>
summarise(
mean = mean(epred),
lo_90 = quantile2(epred, 0.05),
hi_90 = quantile2(epred, 0.95),
.groups = "drop"
) |>
mutate(across(where(is.numeric), ~round(., 2)))# A tibble: 10 × 5
coef vowel mean lo_90 hi_90
<chr> <chr> <dbl> <dbl> <dbl>
1 a a 0.11 -0.14 0.36
2 a e 0.18 -0.02 0.37
3 a i -0.65 -0.74 -0.55
4 a o 0.29 0.17 0.4
5 a u -0.31 -0.44 -0.19
6 b a 0.35 0.14 0.56
7 b e 0.44 0.22 0.66
8 b i 0.06 -0.06 0.17
9 b o 0.34 0.14 0.54
10 b u 0 -0.13 0.124.3 Coefficient b
This section focusses on the coefficient b, the effect of F1, by vowel and place of articulation.
bm_1_draws_long_avg_pl_vow <- bm_1_draws_long |>
group_by(.chain, .iteration, .draw, coef, vowel, c1_place, c2_place) |>
summarise(epred = mean(value), .groups = "drop") |>
mutate(
c1_place = str_remove(c1_place, "c1_place") |> str_sub(1, 3),
c2_place = str_remove(c2_place, "c2_place") |> str_sub(1, 3),
vowel = str_remove(vowel, "vowel")
) |>
unite("c1_c2", c1_place, c2_place)
bm_1_draws_long_avg_pl_vow |>
filter(coef == "b") |>
group_by(c1_c2, vowel) |>
summarise(
mean = mean(epred), sd = sd(epred),
lo_90 = quantile2(epred, 0.05),
hi_90 = quantile2(epred, 0.95)
) |>
mutate(
across(where(is.numeric), ~round(., 2))
)`summarise()` has grouped output by 'c1_c2'. You can override using the
`.groups` argument.# A tibble: 45 × 6
# Groups: c1_c2 [9]
c1_c2 vowel mean sd lo_90 hi_90
<chr> <chr> <dbl> <dbl> <dbl> <dbl>
1 cor_cor a 0.35 0.17 0.07 0.63
2 cor_cor e 0.42 0.25 0.01 0.83
3 cor_cor i 0.08 0.21 -0.26 0.44
4 cor_cor o 0.73 0.24 0.33 1.12
5 cor_cor u 0.02 0.22 -0.33 0.37
6 cor_lab a 0.5 0.15 0.27 0.75
7 cor_lab e 0.58 0.28 0.13 1.03
8 cor_lab i 0.03 0.22 -0.33 0.37
9 cor_lab o 0.3 0.58 -0.66 1.2
10 cor_lab u -0.11 0.19 -0.43 0.2
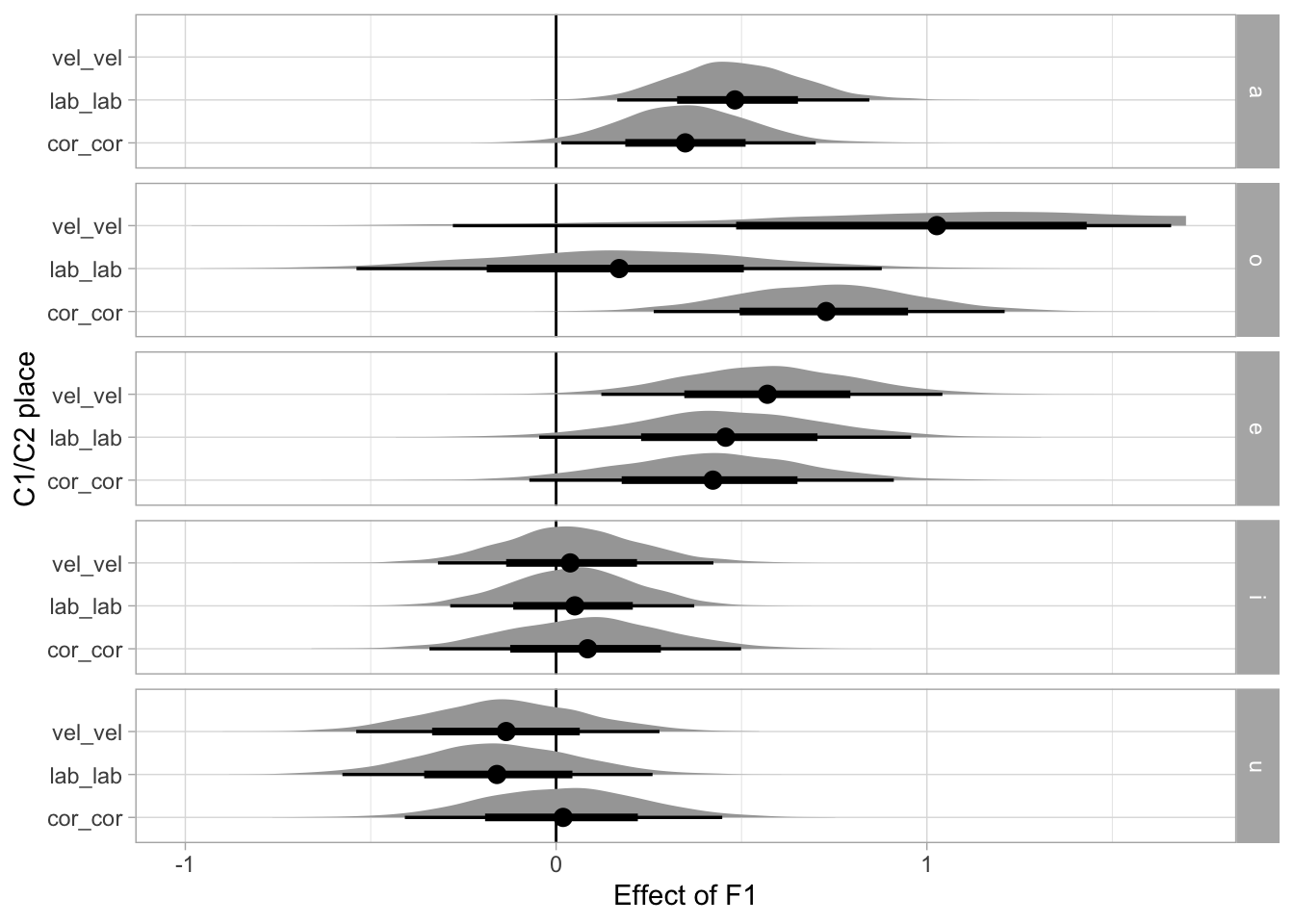
# ℹ 35 more rowsThe following plots the posterior density distributions of the b coefficient by vowel and place of articulation
bm_1_draws_long_avg_pl_vow |>
filter(coef == "b", !(c1_c2 == "lab_cor" & vowel == "e"), !(c1_c2 == "vel_vel" & vowel == "a")) |>
mutate(vowel = factor(vowel, levels = c("a", "o", "e", "i", "u"))) |>
ggplot(aes(epred, c1_c2)) +
geom_vline(xintercept = 0) +
stat_halfeye() +
facet_grid(cols = vars(vowel)) +
xlim(-1, 1.7)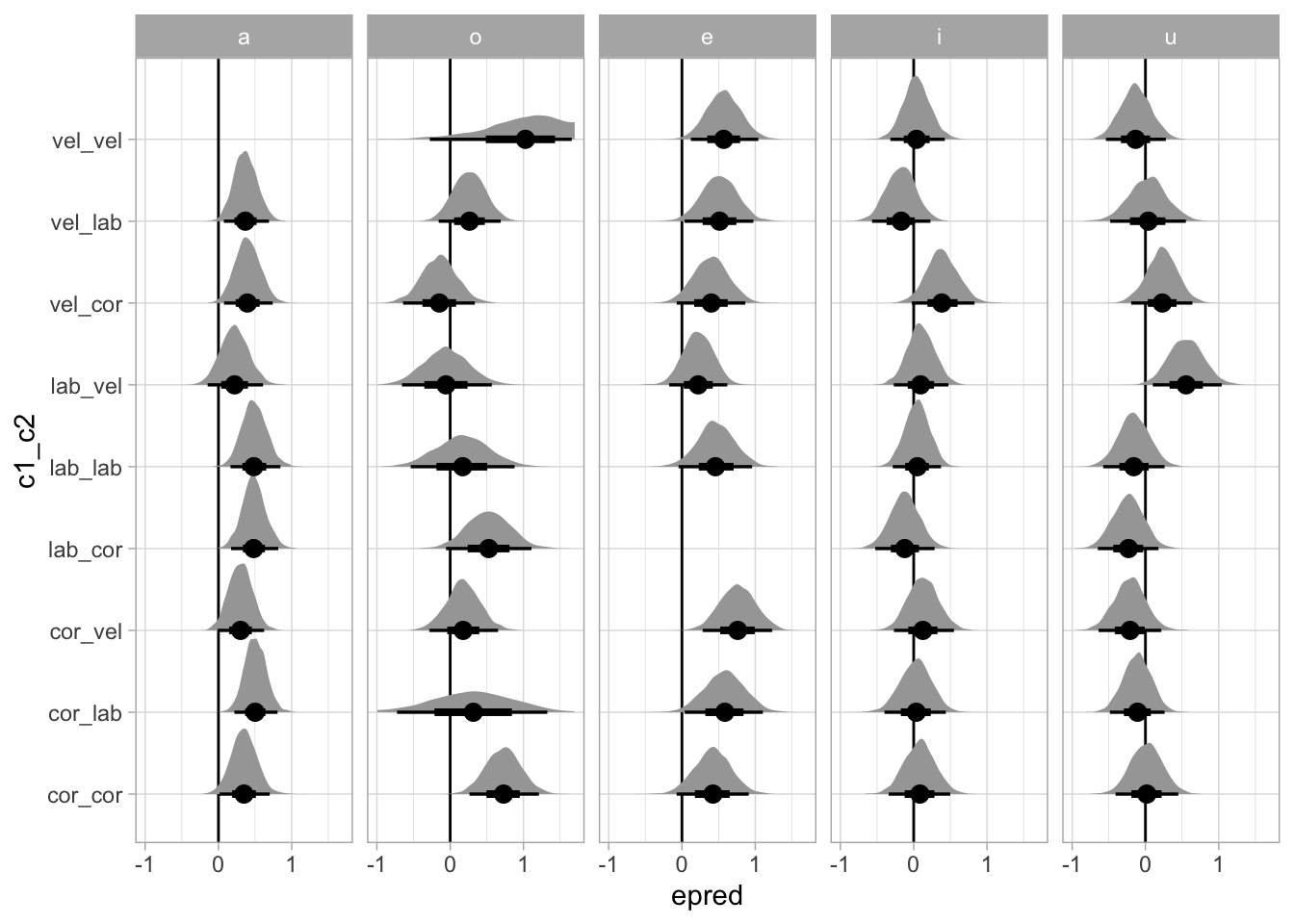
This is the same plot as above but including only homorganic place combinations.
bm_1_draws_long_avg_pl_vow |>
filter(coef == "b", !(c1_c2 == "lab_cor" & vowel == "e"), !(c1_c2 == "vel_vel" & vowel == "a"), c1_c2 %in% c("lab_lab", "cor_cor", "vel_vel")) |>
mutate(vowel = factor(vowel, levels = c("a", "o", "e", "i", "u"))) |>
ggplot(aes(epred, c1_c2)) +
geom_vline(xintercept = 0) +
stat_halfeye() +
facet_grid(rows = vars(vowel)) +
xlim(-1, 1.7) +
labs(
x = "Effect of F1", y = "C1/C2 place"
)
ggsave("img/bm1-f1-place.png", width = 5, height = 5)4.4 Coefficient tables
The following table shows 95, 90, 80 and 60% CrI for the a coefficient by vowel (the intercept by vowel).
bm_1_coef_table_a <- bm_1_draws_long_avg |>
filter(coef == "a") |>
group_by(vowel) |>
reframe(
q95 = round(quantile2(epred, probs = c(0.025, 0.975)), 2),
q90 = round(quantile2(epred, probs = c(0.05, 0.95)), 2),
q80 = round(quantile2(epred, probs = c(0.1, 0.9)), 2),
q60 = round(quantile2(epred, probs = c(0.2, 0.8)), 2)
) |>
mutate(limit = rep(c("lo", "hi"), length.out = n())) |>
pivot_wider(names_from = limit, values_from = q95:q60) |>
unite("q95", q95_lo, q95_hi, sep = ", ") |>
unite("q90", q90_lo, q90_hi, sep = ", ") |>
unite("q80", q80_lo, q80_hi, sep = ", ") |>
unite("q60", q60_lo, q60_hi, sep = ", ")
bm_1_coef_table_a# A tibble: 5 × 5
vowel q95 q90 q80 q60
<chr> <chr> <chr> <chr> <chr>
1 a -0.19, 0.41 -0.14, 0.36 -0.09, 0.31 -0.02, 0.24
2 e -0.06, 0.41 -0.02, 0.37 0.03, 0.33 0.08, 0.28
3 i -0.77, -0.54 -0.74, -0.55 -0.72, -0.58 -0.7, -0.6
4 o 0.15, 0.42 0.17, 0.4 0.2, 0.37 0.23, 0.34
5 u -0.46, -0.16 -0.44, -0.19 -0.41, -0.21 -0.37, -0.25# bm_1_coef_table_a |> knitr::kable(format = "latex") |> cat(sep = "\n")The following table shows 95, 90, 80 and 60% CrI for the b coefficient by vowel and C1/C2 place (the effect of F1).
bm_1_coef_table_b_2 <- bm_1_draws_long_avg_pl_vow |>
filter(coef == "b") |>
group_by(vowel, c1_c2) |>
reframe(
q95 = round(quantile2(epred, probs = c(0.025, 0.975)), 2),
q90 = round(quantile2(epred, probs = c(0.05, 0.95)), 2),
q80 = round(quantile2(epred, probs = c(0.1, 0.9)), 2),
q60 = round(quantile2(epred, probs = c(0.2, 0.8)), 2)
) |>
mutate(limit = rep(c("lo", "hi"), length.out = n())) |>
pivot_wider(names_from = limit, values_from = q95:q60) |>
unite("q95", q95_lo, q95_hi, sep = ", ") |>
unite("q90", q90_lo, q90_hi, sep = ", ") |>
unite("q80", q80_lo, q80_hi, sep = ", ") |>
unite("q60", q60_lo, q60_hi, sep = ", ")
bm_1_coef_table_b_2# A tibble: 45 × 6
vowel c1_c2 q95 q90 q80 q60
<chr> <chr> <chr> <chr> <chr> <chr>
1 a cor_cor 0.01, 0.7 0.07, 0.63 0.13, 0.57 0.2, 0.49
2 a cor_lab 0.22, 0.81 0.27, 0.75 0.32, 0.69 0.38, 0.63
3 a cor_vel -0.01, 0.62 0.04, 0.57 0.09, 0.51 0.16, 0.44
4 a lab_cor 0.17, 0.82 0.22, 0.76 0.28, 0.7 0.35, 0.61
5 a lab_lab 0.17, 0.84 0.22, 0.77 0.27, 0.71 0.34, 0.63
6 a lab_vel -0.15, 0.61 -0.09, 0.55 -0.03, 0.47 0.06, 0.38
7 a vel_cor 0.08, 0.74 0.12, 0.68 0.18, 0.62 0.25, 0.54
8 a vel_lab 0.07, 0.69 0.11, 0.64 0.17, 0.58 0.24, 0.5
9 a vel_vel -1.97, 1.94 -1.66, 1.65 -1.27, 1.29 -0.84, 0.83
10 e cor_cor -0.07, 0.91 0.01, 0.83 0.1, 0.74 0.21, 0.63
# ℹ 35 more rows# bm_1_coef_table_b_2 |> knitr::kable(format = "latex") |> cat(sep = "\n")4.5 Posterior predictive check and sensitivity analysis
Posterior predictive checks are good.
pp_check(bm_1, ndraws = 50)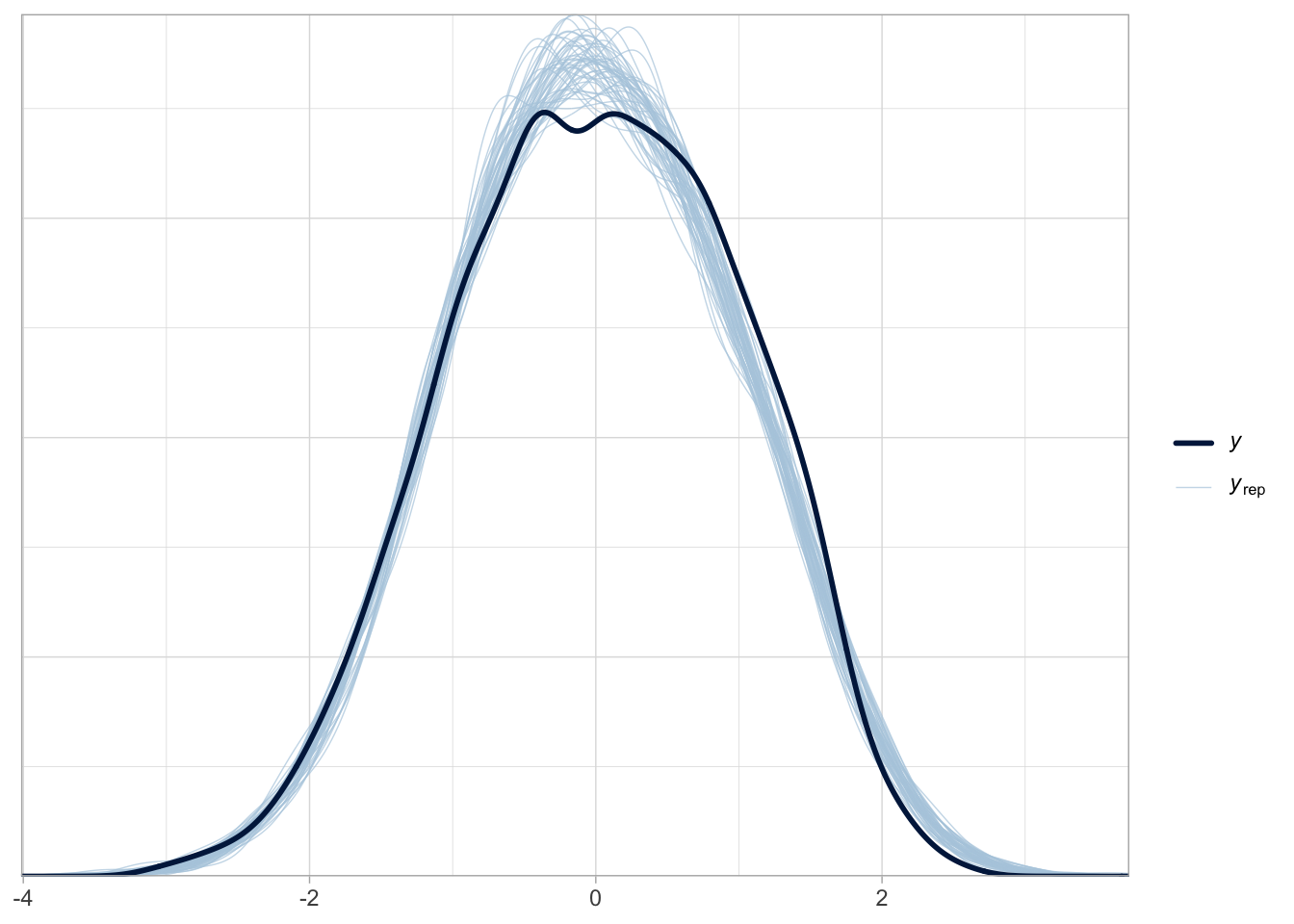
bm_1_fix <- fixef(bm_1) |> as_tibble(rownames = "term") |>
mutate(
theta = rep(0, 91),
sigma_prior = c(rep(1, 90), 0.1),
z = abs((Estimate - theta) / Est.Error),
s = 1 - (Est.Error^2 / sigma_prior^2),
term = str_remove(term, "vowel") |> str_remove_all("(c1|c2)_place")
)labels <- tibble(
s = c(0.25, 0.25, 0.75, 0.75),
z = c(1.25, 3.75, 1.25, 3.75),
term = c("Poorly identified", "Prior/Posterior\nconflict", "Ideal", "Overfit")
)
bm_1_fix |>
ggplot(aes(s, z, label = term)) +
annotate("rect", xmin = 0, ymin = 0, xmax = 0.5, ymax = 2.5, alpha = 0.5, fill = "#e66101") +
annotate("rect", xmin = 0, ymin = 2.5, xmax = 0.5, ymax = Inf, alpha = 0.5, fill = "#fdb863") +
annotate("rect", xmin = 0.5, ymin = 0, xmax = 1, ymax = 2.5, alpha = 0.5, fill = "#b2abd2") +
annotate("rect", xmin = 0.5, ymin = 2.5, xmax = 1, ymax = Inf, alpha = 0.5, fill = "#5e3c99") +
geom_text(size = 3, alpha = 0.5) +
# geom_point(alpha = 0.5) +
geom_label(data = labels, colour = "white", fill = "black") +
xlim(0, 1)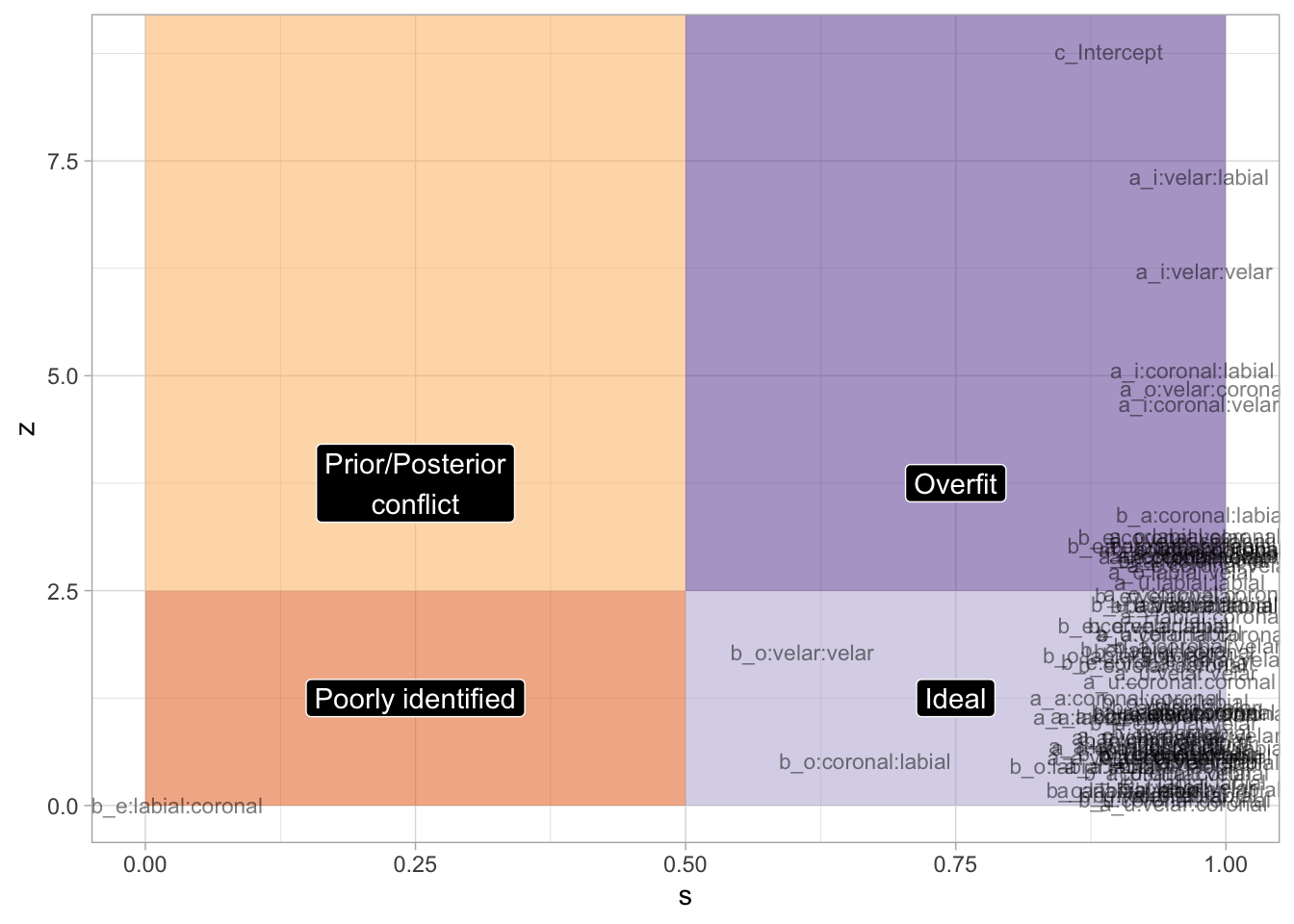
4.6 Model plotting
We need to get the predicted draws to convert duration and F1 back to ms and hz. Note that duration was logged then scaled.
seq_minmax <- function(x, by = 1) {
seq(min(x), max(x), by = by)
}
bm_1_preds <- bm_1_draws_long |>
group_by(coef, vowel, .draw) |>
summarise(
value = mean(value),
.groups = "drop"
) |>
pivot_wider(names_from = coef, values_from = value) |>
mutate(
f13_z = list(seq_minmax(ita_egg$f13_z, 0.5))
) |>
unnest_longer(f13_z) |>
mutate(
epred = a + b * f13_z,
duration_log = epred * sd(ita_egg$duration_log) + mean(ita_egg$duration_log),
duration = exp(duration_log),
f13 = f13_z * sd(ita_egg$f13) + mean(ita_egg$f13),
vowel = str_remove(vowel, "vowel")
)Let’s also calculate the mean F1 values for each vowel, to be added in the plot below.
vmean_f13 <- ita_egg |>
group_by(vowel) |>
summarise(f13_mean = mean(f13))
vmean_f13z <- ita_egg |>
group_by(vowel) |>
summarise(f13z_mean = mean(f13_z))We can now plot the model predictions in the original scale.
bm_1_preds |>
group_by(vowel, f13) |>
summarise(
duration_ms = median(duration),
# Get the 90% CrI
q0.05 = quantile(duration, probs = 0.05),
q0.90 = quantile(duration, probs = 0.95),
.groups = "drop"
) |>
ggplot(aes(f13, duration_ms)) +
geom_ribbon(aes(ymin = q0.05, ymax = q0.90, fill = vowel), alpha = 0.25) +
geom_line(aes(colour = vowel), linewidth = 1) +
facet_wrap(~vowel) +
scale_colour_viridis_d() +
scale_fill_viridis_d()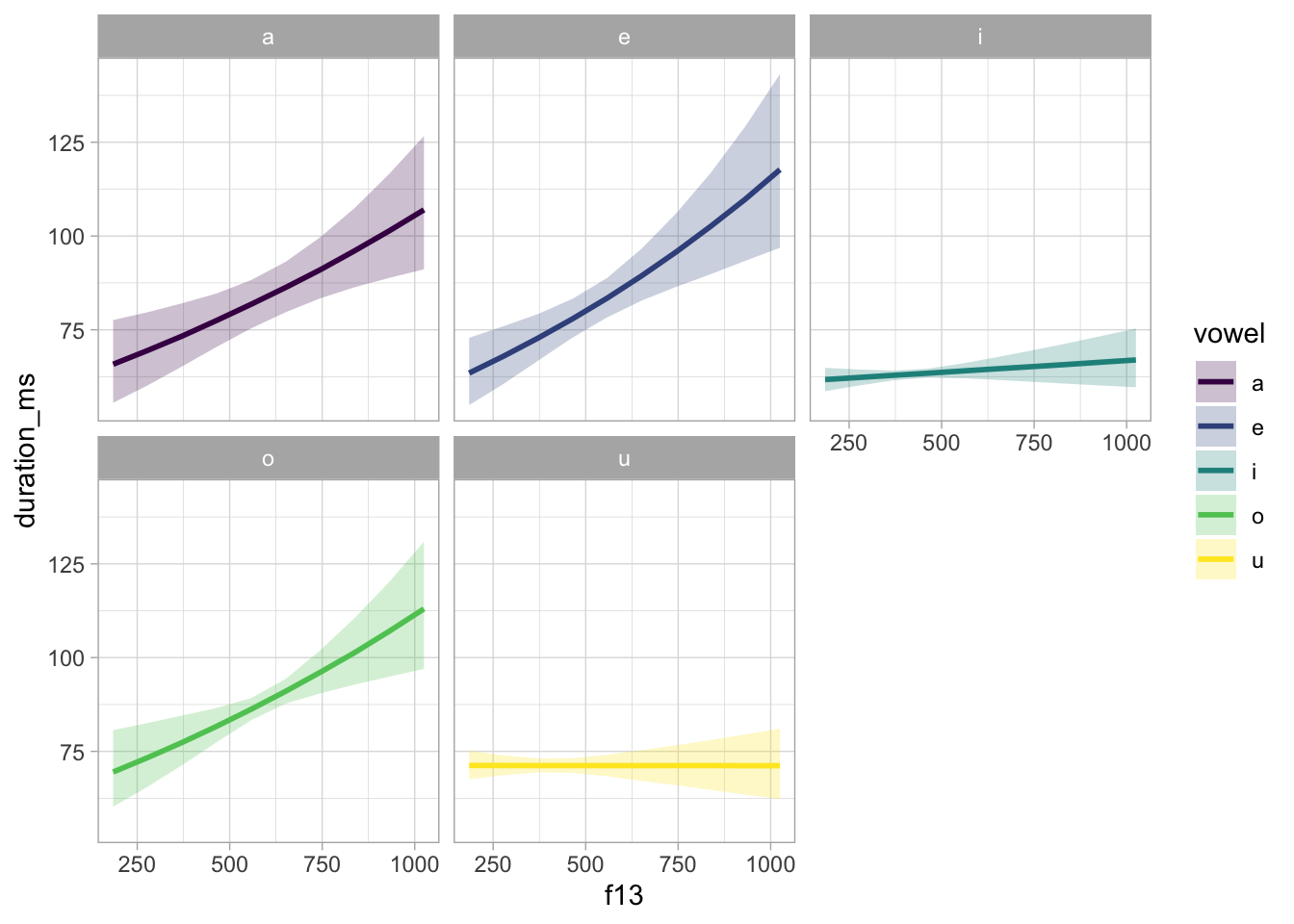
bm_1_preds |>
group_by(vowel, f13) |>
ggplot(aes(f13, duration)) +
stat_lineribbon() +
facet_wrap(~vowel) +
scale_fill_brewer()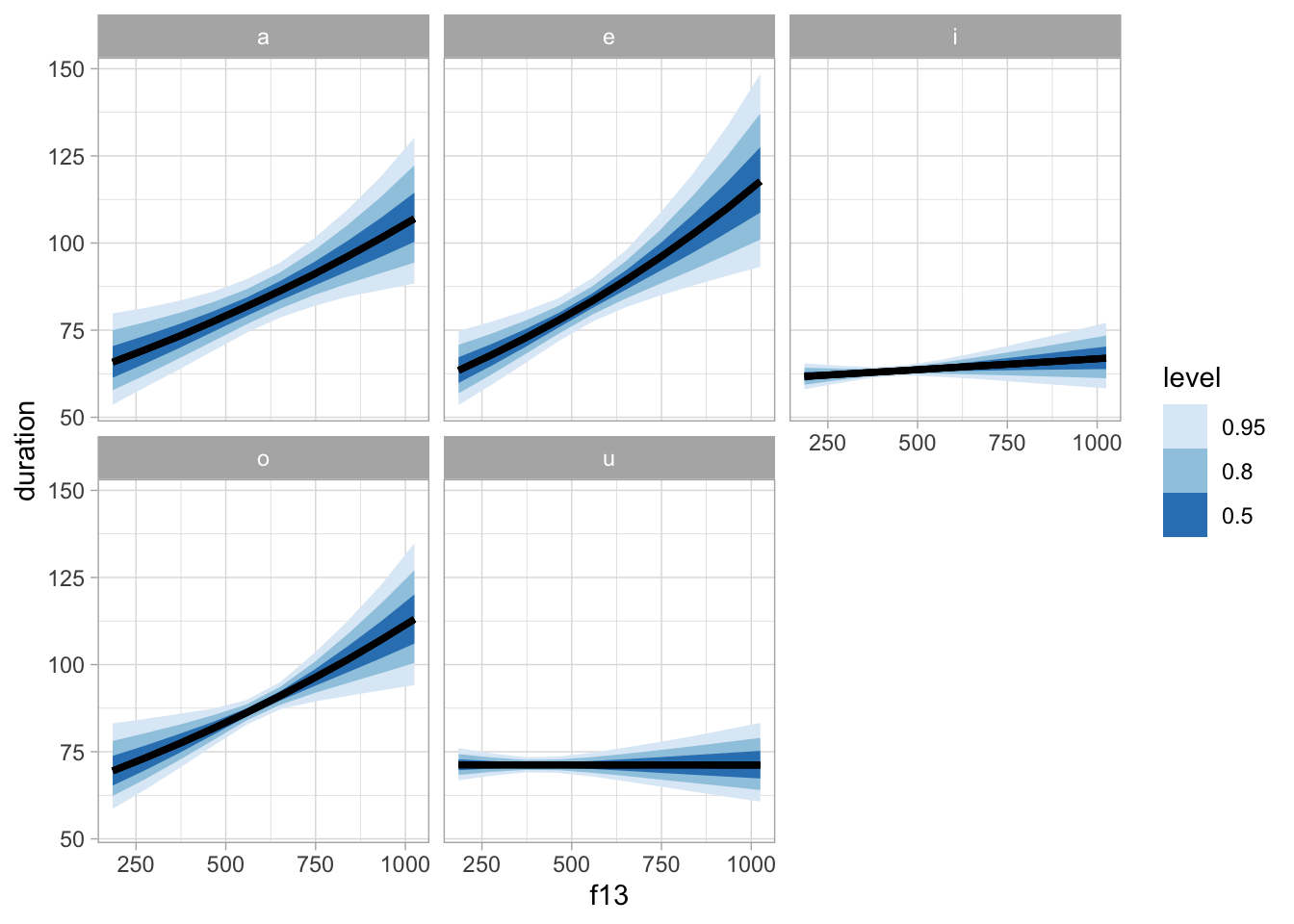
bm_1_preds |>
mutate(
vowel = str_replace(vowel, "o", "ɔ"),
vowel = factor(vowel, levels = c("i", "u", "e", "ɔ", "a"))
) |>
group_by(vowel, f13) |>
ggplot(aes(f13, duration, fill = vowel)) +
stat_ribbon(.width = 0.98, alpha = 0.2) +
stat_ribbon(.width = 0.9, alpha = 0.4) +
stat_lineribbon(.width = 0.6, alpha = 0.8) +
geom_vline(data = vmean_f13 |> mutate(
vowel = str_replace(vowel, "o", "ɔ"),
vowel = factor(vowel, levels = c("i", "u", "e", "ɔ", "a"))
), aes(xintercept = f13_mean, colour = vowel), linetype = "dashed") +
geom_rug(data = ita_egg |> mutate(
vowel = str_replace(vowel, "o", "ɔ"),
vowel = factor(vowel, levels = c("i", "u", "e", "ɔ", "a"))
), alpha = 0.1, length = unit(0.015, "npc"), aes(colour = vowel)) +
facet_wrap(~vowel) +
labs(
x = "F1 (Hz)", y = "Duration (ms)"
) +
scale_fill_manual(values = cols) +
scale_color_manual(values = cols) +
theme(legend.position = "none")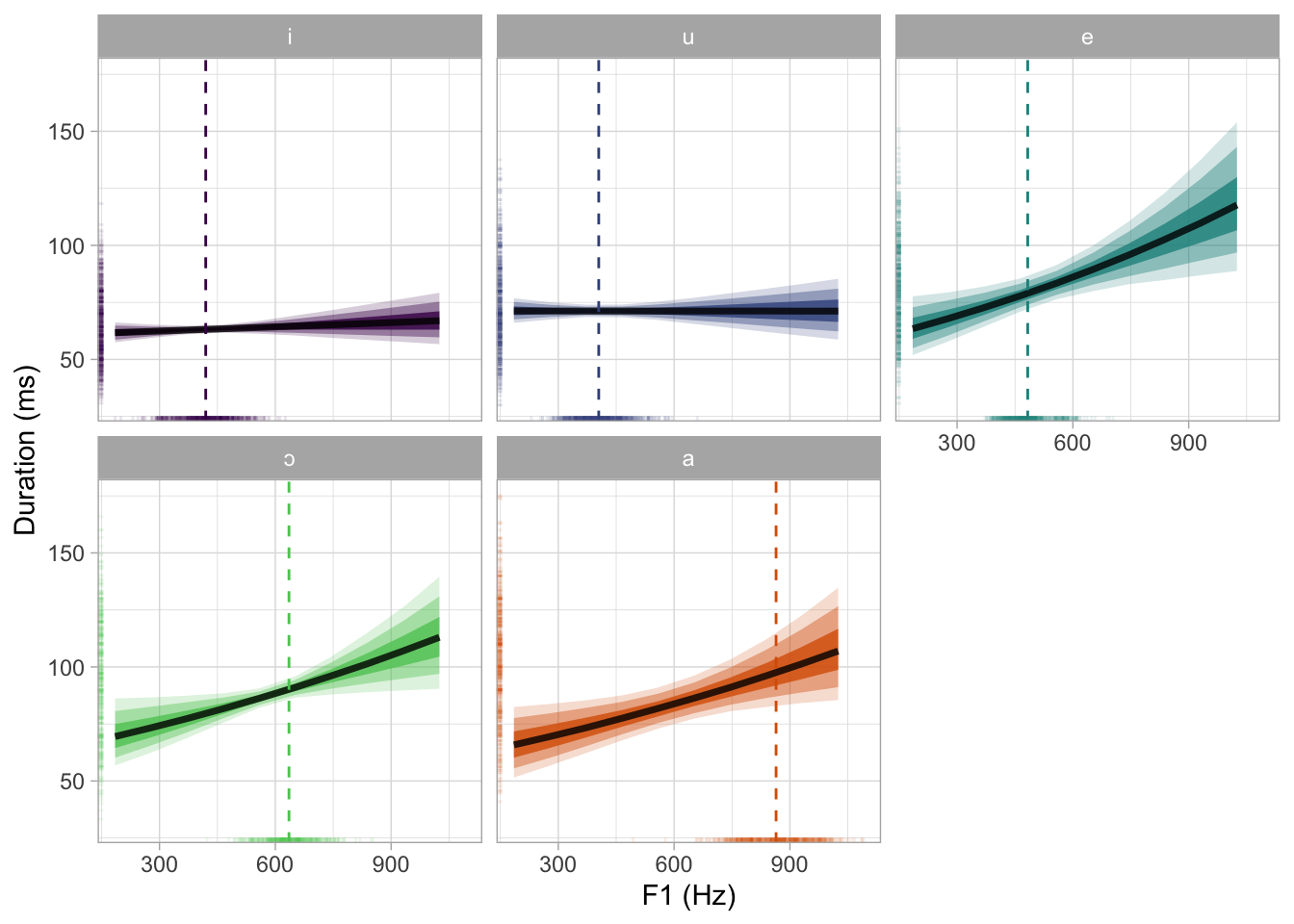
ggsave("img/bm1-pred-plot-ms-hz.png", width = 7, height = 5)But let’s also plot this in the standardised logged duration scale.
bm_1_preds |>
group_by(vowel, f13) |>
ggplot(aes(f13_z, epred, fill = vowel)) +
stat_ribbon(.width = 0.98, alpha = 0.2) +
stat_ribbon(.width = 0.9, alpha = 0.4) +
stat_lineribbon(.width = 0.6, alpha = 0.8) +
geom_vline(data = vmean_f13z, aes(xintercept = f13z_mean, colour = vowel)) +
facet_wrap(~vowel) +
scale_fill_viridis_d()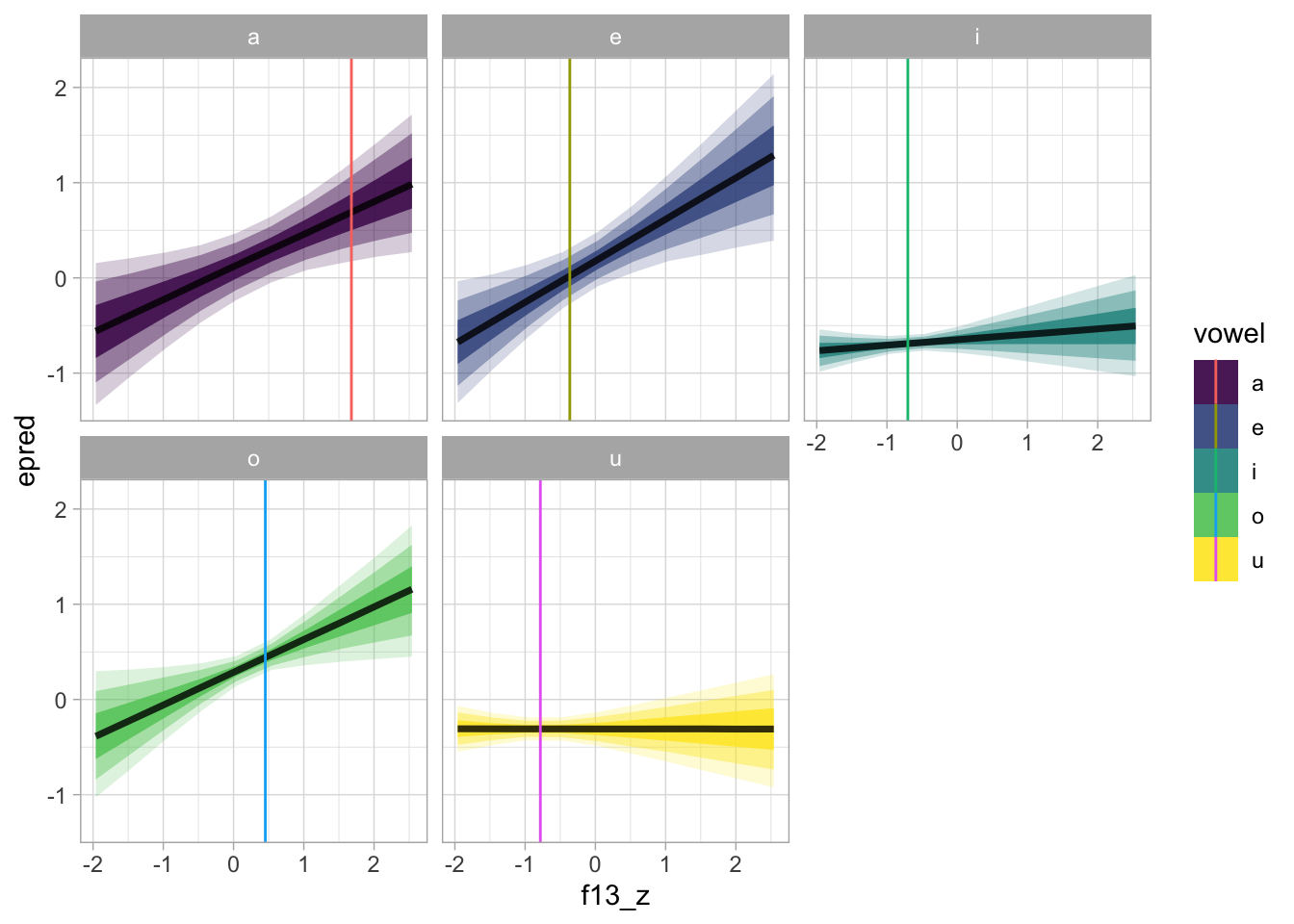
4.7 Group-level effects
bm_1_ranef <- bm_1 |>
gather_draws(r_speaker__c[subject,var])bm_1_ranef |>
ggplot(aes(y = reorder(subject, .value, median), x = .value)) +
geom_vline(xintercept = 0) +
stat_halfeye()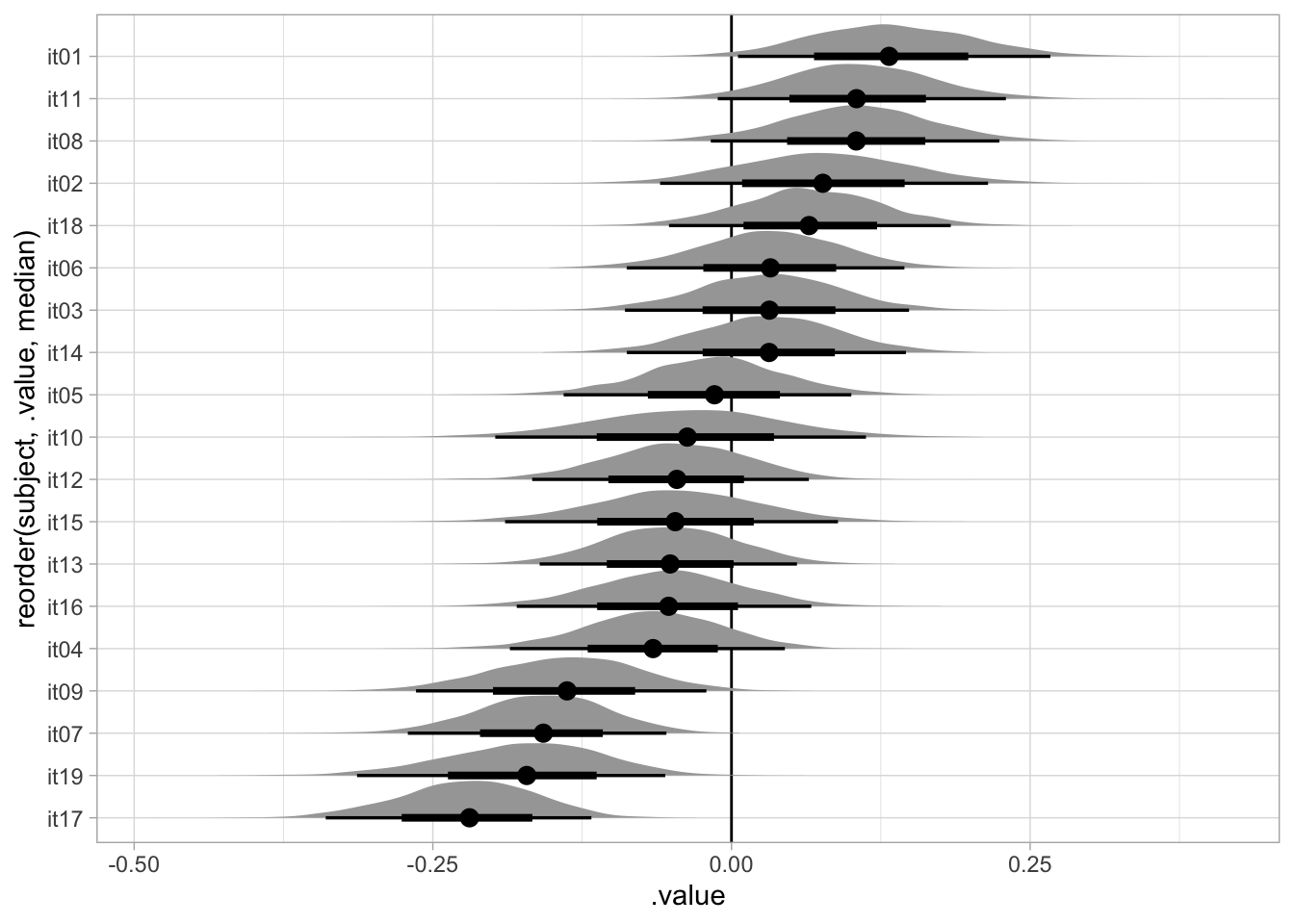
5 Non-linear modelling: GAM
A GAM model with the mgcv package is run to get an estimate number of k since brms does not optimise k and fits the model just with the default k which can slow down the MCMC sampling with large values of k.
gam_1 <- bam(
duration_logz ~
vowel +
s(f13_z) +
s(f13_z, speaker, by = vowel, bs = "fs", m = 1) +
s(speech_rate_logz) +
s(speech_rate_logz, speaker, bs = "fs", m = 1),
data = ita_egg
)Warning in gam.side(sm, X, tol = .Machine$double.eps^0.5): model has repeated
1-d smooths of same variable.summary(gam_1)
Family: gaussian
Link function: identity
Formula:
duration_logz ~ vowel + s(f13_z) + s(f13_z, speaker, by = vowel,
bs = "fs", m = 1) + s(speech_rate_logz) + s(speech_rate_logz,
speaker, bs = "fs", m = 1)
Parametric coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.10762 0.14848 0.725 0.4686
vowele 0.05915 0.14477 0.409 0.6829
voweli -0.59166 0.15070 -3.926 8.83e-05 ***
vowelo 0.28281 0.12060 2.345 0.0191 *
vowelu -0.22071 0.16073 -1.373 0.1698
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Approximate significance of smooth terms:
edf Ref.df F p-value
s(f13_z) 3.484 4.235 12.819 < 2e-16 ***
s(f13_z,speaker):vowela 16.684 144.000 0.346 < 2e-16 ***
s(f13_z,speaker):vowele 17.048 137.000 0.247 1.19e-06 ***
s(f13_z,speaker):voweli 11.431 148.000 0.121 3.31e-05 ***
s(f13_z,speaker):vowelo 18.291 147.000 0.315 < 2e-16 ***
s(f13_z,speaker):vowelu 24.993 151.000 0.606 < 2e-16 ***
s(speech_rate_logz) 4.183 4.948 20.333 < 2e-16 ***
s(speech_rate_logz,speaker) 52.949 170.000 2.523 < 2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
R-sq.(adj) = 0.742 Deviance explained = 75.5%
fREML = 2415.4 Scale est. = 0.2583 n = 3019vmean <- aggregate(ita_egg$f13_z, list(ita_egg$vowel), mean)
# fs_terms <- c("s(f13_z,speaker)")
fs_terms <- c("s(f13_z,speaker):vowela", "s(f13_z,speaker):vowele", "s(f13_z,speaker):voweli", "s(f13_z,speaker):vowelo", "s(f13_z,speaker):vowelu", "s(speech_rate_logz,speaker)")
predict_gam(gam_1, exclude_terms = fs_terms, length_out = 100, values = list(speech_rate_logz = 0)) |>
plot(series = "f13_z", comparison = "vowel") +
geom_vline(data = vmean, aes(xintercept = x, colour = Group.1)) +
geom_hline(yintercept = 0, linetype = "dashed") +
facet_wrap(~vowel)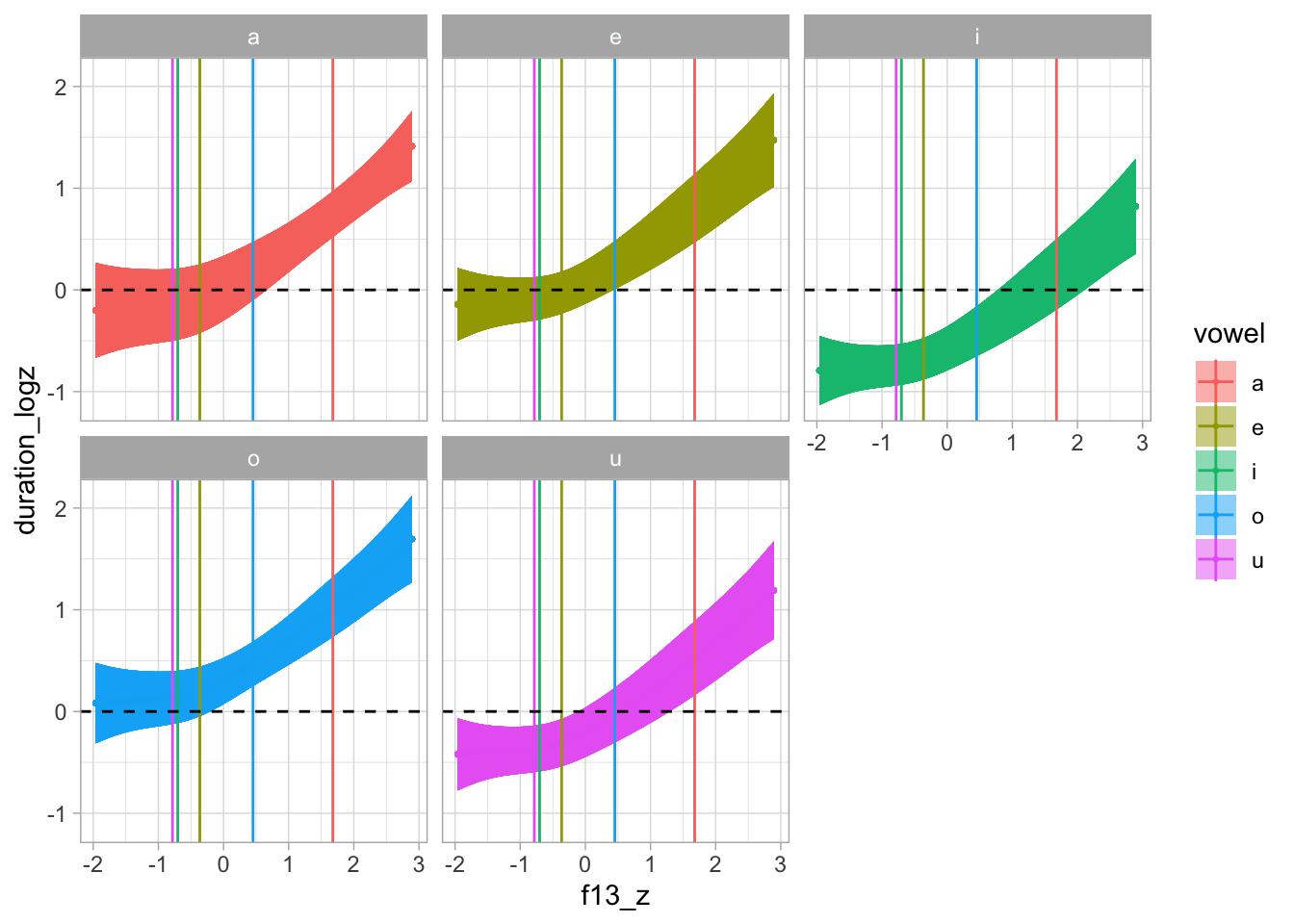
6 Non-linear modelling: BRM
6.1 Prior predictive checks
priors_s <- c(
prior(normal(0, 1), class = b),
prior(cauchy(0, 0.01), class = sigma),
prior(cauchy(0, 1), class = sds)
)
bms_1_f <- bf(
duration_logz ~
0 + vowel +
s(f13_z, k = 5) +
s(f13_z, speaker, by = vowel, bs = "fs", m = 1) +
s(speech_rate_logz, k = 5) +
s(speech_rate_logz, speaker, bs = "fs", m = 1)
)
bms_1_priors <- brm(
bms_1_f,
family = gaussian,
data = ita_egg,
prior = priors_s,
sample_prior = "only",
cores = 4,
threads = threading(2),
backend = "cmdstanr",
file = "data/cache/bms_1_priors",
seed = my_seed
)plot(conditional_smooths(bms_1_priors, ndraws = 100), ask = FALSE)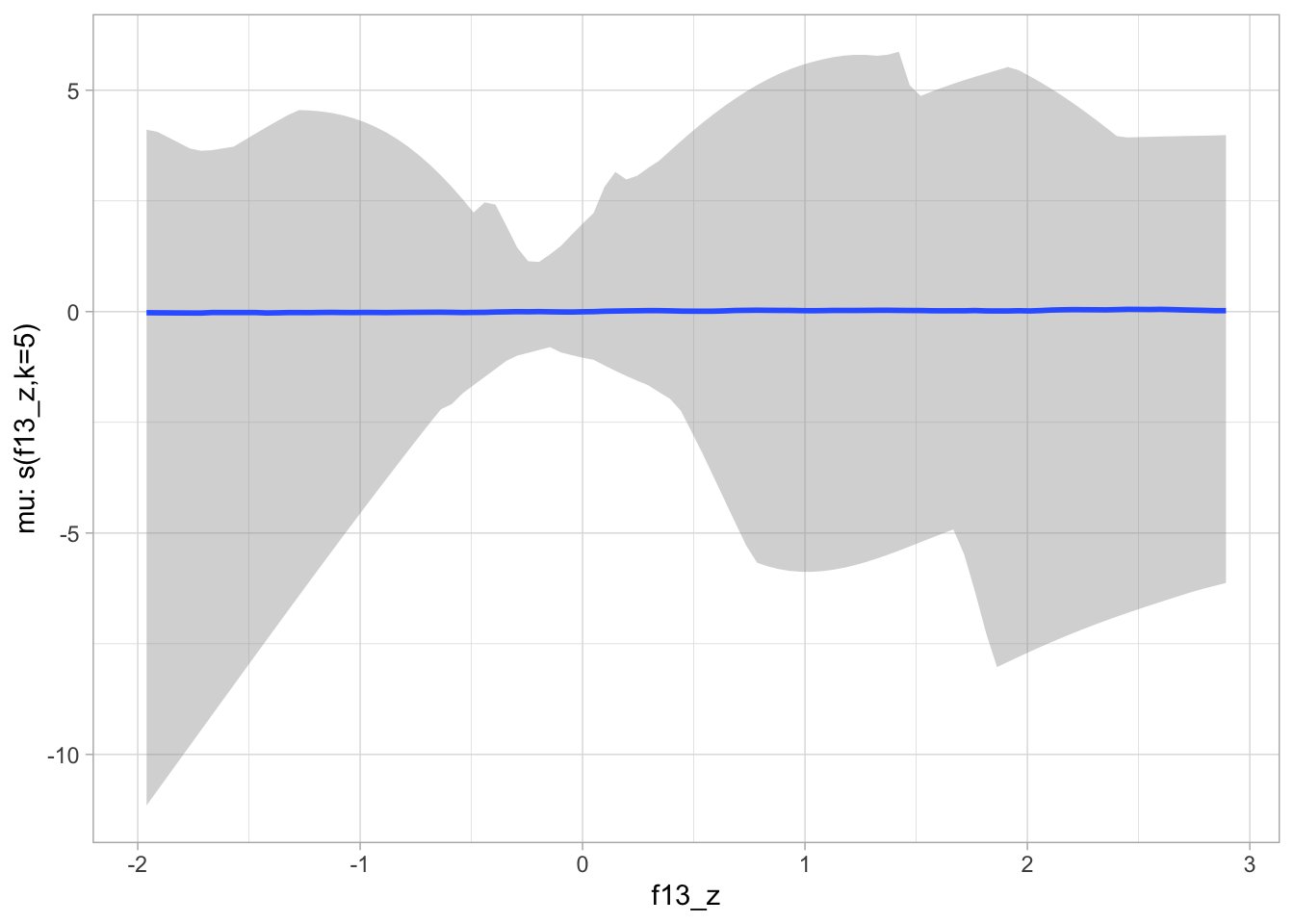
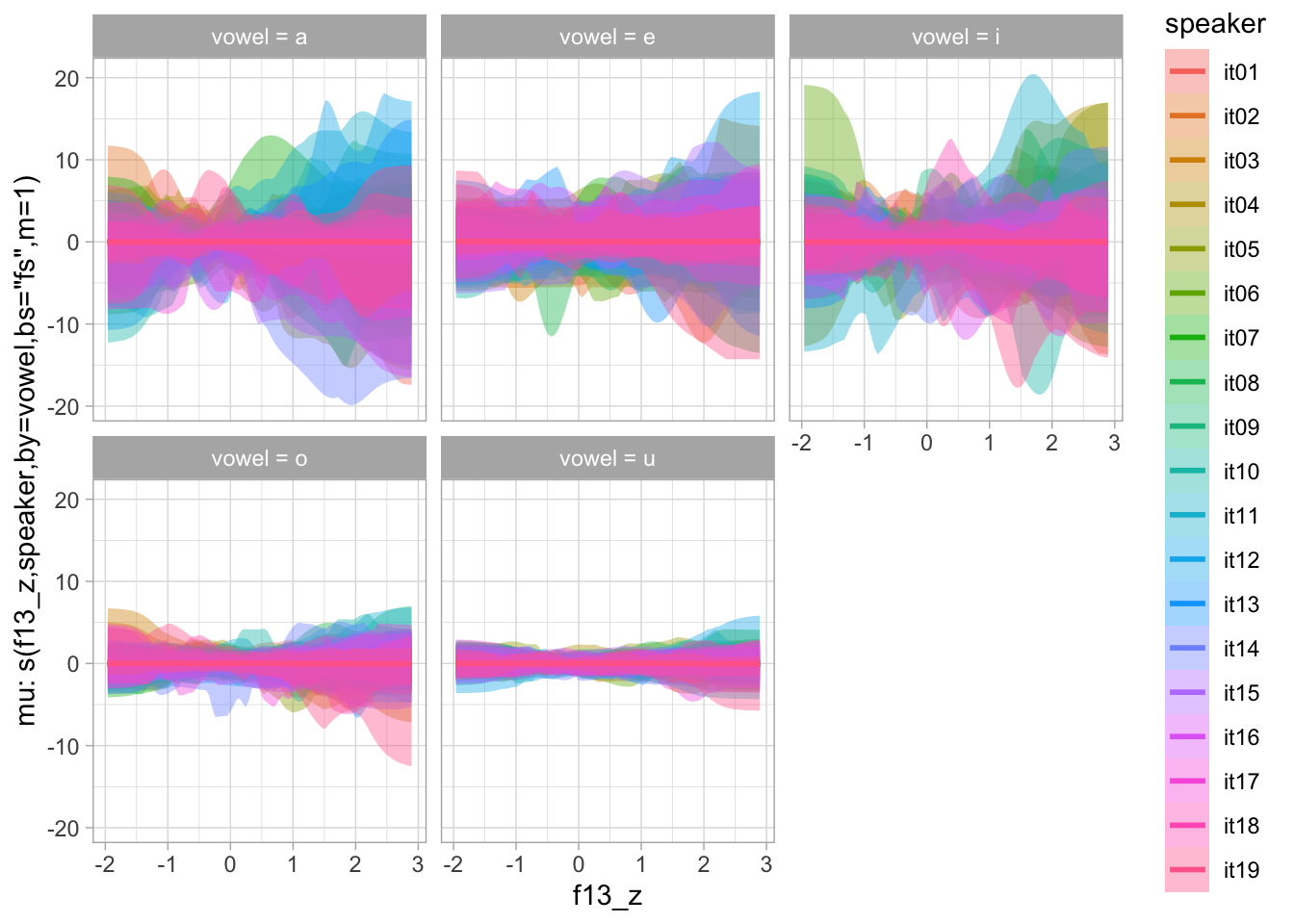
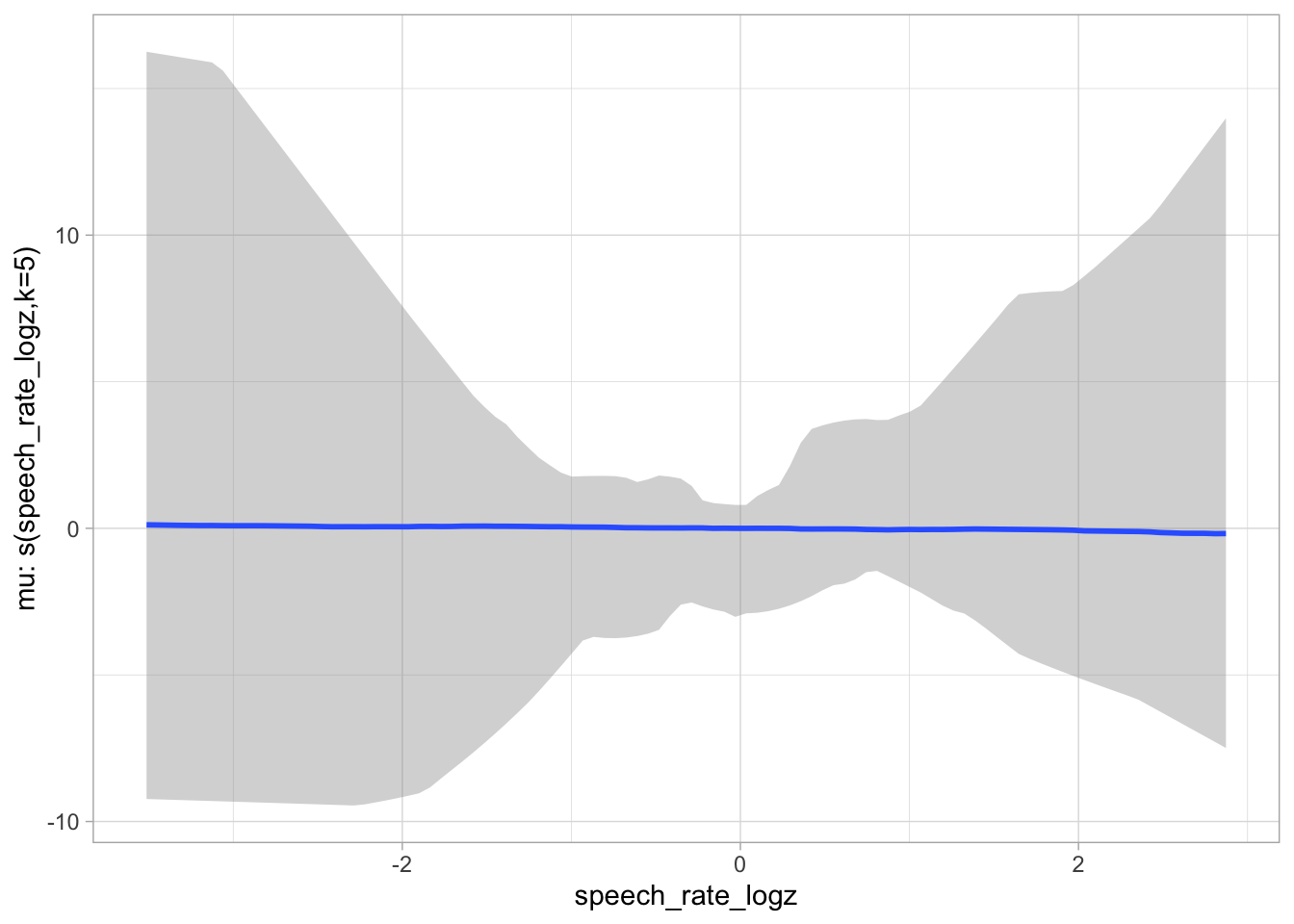
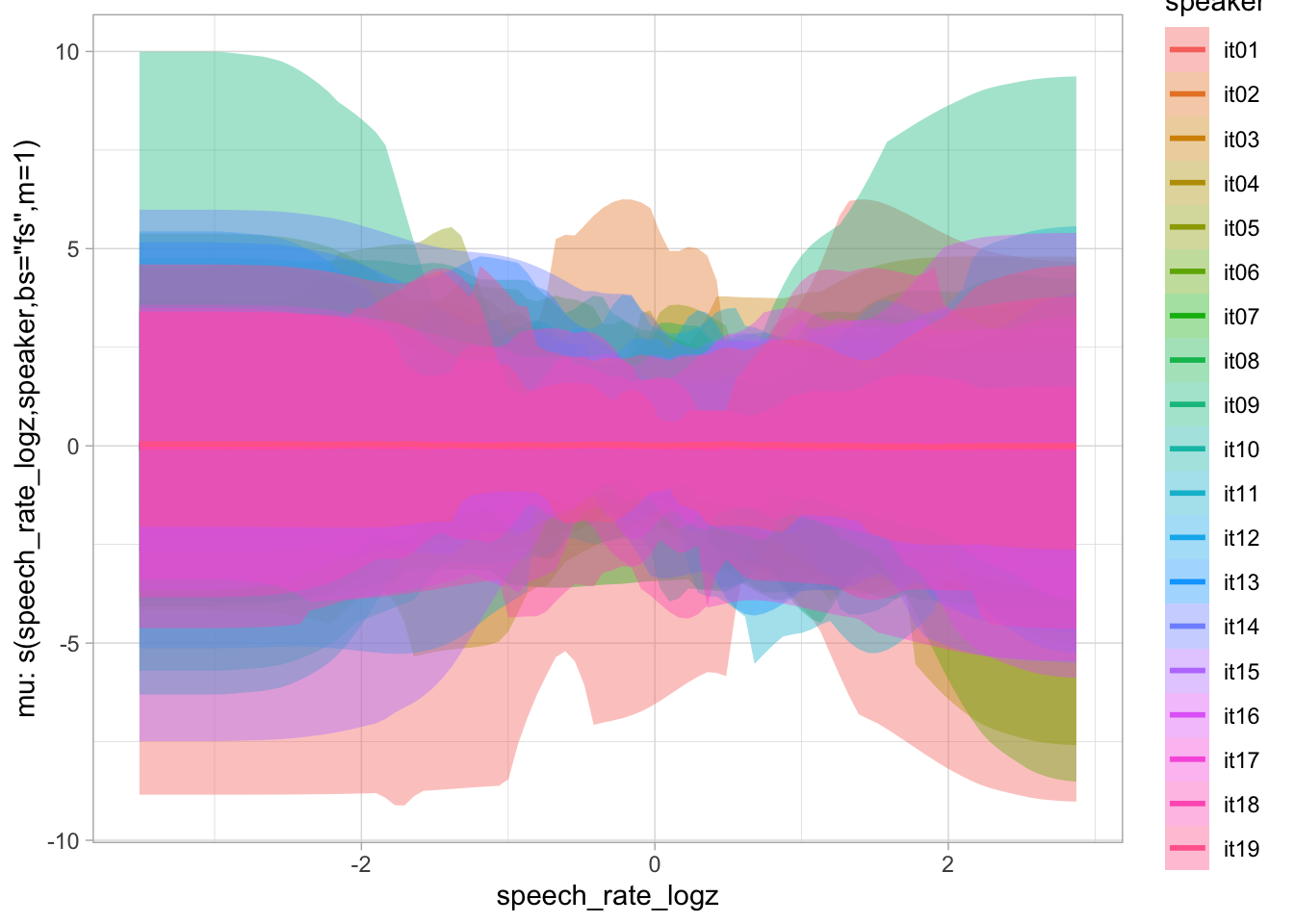
6.2 Model fit
We specify k = 5 based on the mgcv modelling above. Reducing k speeds up estimation (because there are less basis functions, hence less parameters to estimate).
The model takes about 4-5 hours to run on 8 cores.
bms_1 <- brm(
bms_1_f,
family = gaussian,
data = ita_egg,
prior = priors_s,
cores = 4,
threads = threading(2),
backend = "cmdstanr",
file = "data/cache/bms_1",
seed = my_seed
)summary(bms_1, prob = 0.9) Family: gaussian
Links: mu = identity; sigma = identity
Formula: duration_logz ~ 0 + vowel + s(f13_z, k = 5) + s(f13_z, speaker, by = vowel, bs = "fs", m = 1) + s(speech_rate_logz, k = 5) + s(speech_rate_logz, speaker, bs = "fs", m = 1)
Data: ita_egg (Number of observations: 3019)
Draws: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
total post-warmup draws = 4000
Smoothing Spline Hyperparameters:
Estimate Est.Error l-90% CI u-90% CI Rhat
sds(sf13_z_1) 1.57 0.87 0.69 3.22 1.00
sds(sf13_zspeakervowela_1) 0.34 0.23 0.03 0.76 1.00
sds(sf13_zspeakervowela_2) 1.90 1.96 0.08 6.11 1.01
sds(sf13_zspeakervowele_1) 0.33 0.22 0.03 0.74 1.00
sds(sf13_zspeakervowele_2) 1.93 1.96 0.09 6.17 1.00
sds(sf13_zspeakervoweli_1) 0.33 0.22 0.02 0.74 1.01
sds(sf13_zspeakervoweli_2) 1.99 2.03 0.09 6.40 1.00
sds(sf13_zspeakervowelo_1) 0.34 0.23 0.03 0.75 1.01
sds(sf13_zspeakervowelo_2) 1.82 1.90 0.09 5.93 1.00
sds(sf13_zspeakervowelu_1) 0.32 0.23 0.03 0.74 1.01
sds(sf13_zspeakervowelu_2) 2.04 2.03 0.08 6.17 1.01
sds(sspeech_rate_logz_1) 0.45 0.44 0.03 1.23 1.00
sds(sspeech_rate_logzspeaker_1) 0.78 0.13 0.58 1.00 1.00
sds(sspeech_rate_logzspeaker_2) 1.86 1.93 0.08 5.95 1.01
Bulk_ESS Tail_ESS
sds(sf13_z_1) 2051 2236
sds(sf13_zspeakervowela_1) 492 1166
sds(sf13_zspeakervowela_2) 721 1055
sds(sf13_zspeakervowele_1) 562 884
sds(sf13_zspeakervowele_2) 818 1027
sds(sf13_zspeakervoweli_1) 404 1003
sds(sf13_zspeakervoweli_2) 607 802
sds(sf13_zspeakervowelo_1) 336 990
sds(sf13_zspeakervowelo_2) 581 909
sds(sf13_zspeakervowelu_1) 509 890
sds(sf13_zspeakervowelu_2) 619 1091
sds(sspeech_rate_logz_1) 1715 2314
sds(sspeech_rate_logzspeaker_1) 1573 2465
sds(sspeech_rate_logzspeaker_2) 825 1429
Regression Coefficients:
Estimate Est.Error l-90% CI u-90% CI Rhat Bulk_ESS Tail_ESS
vowela 0.08 0.15 -0.16 0.32 1.00 2140 2459
vowele 0.23 0.11 0.05 0.42 1.00 1266 1664
voweli -0.46 0.11 -0.64 -0.28 1.00 1300 1635
vowelo 0.39 0.11 0.20 0.57 1.00 1299 1750
vowelu -0.10 0.11 -0.28 0.08 1.00 1271 1729
sf13_z_1 0.97 0.62 -0.04 1.98 1.00 3746 3388
sspeech_rate_logz_1 -2.57 0.42 -3.25 -1.89 1.00 2522 2503
Further Distributional Parameters:
Estimate Est.Error l-90% CI u-90% CI Rhat Bulk_ESS Tail_ESS
sigma 0.52 0.01 0.51 0.53 1.00 5481 2619
Draws were sampled using sample(hmc). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).bms_1_coef_table <- bms_1 |>
as_draws_df() |>
select(b_vowela:b_vowelu) |>
pivot_longer(b_vowela:b_vowelu) |>
group_by(name) |>
median_hdi()Warning: Dropping 'draws_df' class as required metadata was removed.bms_1_coef_table# A tibble: 5 × 7
name value .lower .upper .width .point .interval
<chr> <dbl> <dbl> <dbl> <dbl> <chr> <chr>
1 b_vowela 0.0797 -0.197 0.368 0.95 median hdi
2 b_vowele 0.234 0.0219 0.457 0.95 median hdi
3 b_voweli -0.459 -0.678 -0.241 0.95 median hdi
4 b_vowelo 0.386 0.175 0.612 0.95 median hdi
5 b_vowelu -0.100 -0.306 0.131 0.95 median hdi # bms_1_coef_table |> knitr::kable(format = "latex") |> cat(sep = "\n")bms_1_coef_table_2 <- bms_1 |>
as_draws_df() |>
select(b_vowela:b_vowelu) |>
pivot_longer(b_vowela:b_vowelu) |>
group_by(name) |>
reframe(
q95 = round(quantile2(value, probs = c(0.025, 0.975)), 2),
q90 = round(quantile2(value, probs = c(0.05, 0.95)), 2),
q80 = round(quantile2(value, probs = c(0.1, 0.9)), 2),
q60 = round(quantile2(value, probs = c(0.2, 0.8)), 2)
) |>
mutate(limit = rep(c("lo", "hi"), length.out = n())) |>
pivot_wider(names_from = limit, values_from = q95:q60) |>
unite("q95", q95_lo, q95_hi, sep = ", ") |>
unite("q90", q90_lo, q90_hi, sep = ", ") |>
unite("q80", q80_lo, q80_hi, sep = ", ") |>
unite("q60", q60_lo, q60_hi, sep = ", ")Warning: Dropping 'draws_df' class as required metadata was removed.bms_1_coef_table_2# A tibble: 5 × 5
name q95 q90 q80 q60
<chr> <chr> <chr> <chr> <chr>
1 b_vowela -0.22, 0.36 -0.16, 0.32 -0.11, 0.27 -0.04, 0.2
2 b_vowele 0.02, 0.45 0.05, 0.42 0.09, 0.37 0.14, 0.32
3 b_voweli -0.68, -0.24 -0.64, -0.28 -0.6, -0.32 -0.55, -0.37
4 b_vowelo 0.16, 0.61 0.2, 0.57 0.24, 0.53 0.3, 0.48
5 b_vowelu -0.32, 0.12 -0.28, 0.08 -0.24, 0.04 -0.19, -0.01# bms_1_coef_table_2 |> knitr::kable(format = "latex") |> cat(sep = "\n")6.3 Posterior predictive check and sensitivity analysis
pp_check(bms_1, ndraws = 50)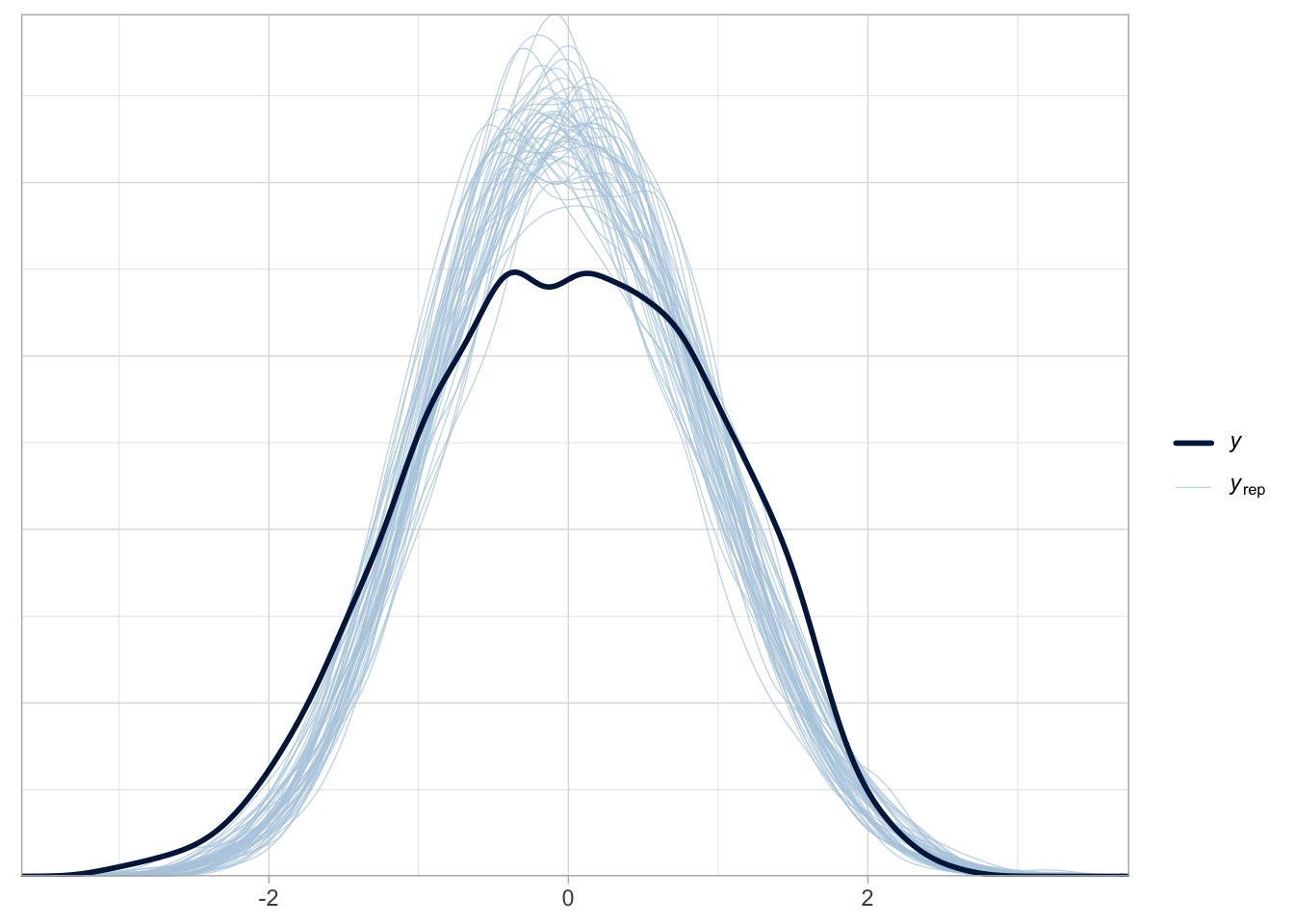
bms_1_fix <- fixef(bms_1) |> as_tibble(rownames = "term") |>
mutate(
theta = rep(0, 7),
sigma_prior = rep(1, 7),
z = abs((Estimate - theta) / Est.Error),
s = 1 - (Est.Error^2 / sigma_prior^2)
)
bms_1_fix# A tibble: 7 × 9
term Estimate Est.Error Q2.5 Q97.5 theta sigma_prior z s
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 vowela 0.0797 0.146 -0.215 0.358 0 1 0.547 0.979
2 vowele 0.235 0.110 0.0170 0.454 0 1 2.12 0.988
3 voweli -0.460 0.111 -0.677 -0.240 0 1 4.16 0.988
4 vowelo 0.387 0.111 0.165 0.608 0 1 3.48 0.988
5 vowelu -0.100 0.111 -0.320 0.123 0 1 0.901 0.988
6 sf13_z_1 0.971 0.621 -0.226 2.18 0 1 1.56 0.614
7 sspeech_rate_… -2.57 0.420 -3.41 -1.74 0 1 6.11 0.823labels <- tibble(
s = c(0.25, 0.25, 0.75, 0.75),
z = c(1.25, 3.75, 1.25, 3.75),
term = c("Poorly identified", "Prior/Posterior\nconflict", "Ideal", "Overfit")
)
bms_1_fix |>
ggplot(aes(s, z, label = term)) +
annotate("rect", xmin = 0, ymin = 0, xmax = 0.5, ymax = 2.5, alpha = 0.5, fill = "#e66101") +
annotate("rect", xmin = 0, ymin = 2.5, xmax = 0.5, ymax = Inf, alpha = 0.5, fill = "#fdb863") +
annotate("rect", xmin = 0.5, ymin = 0, xmax = 1, ymax = 2.5, alpha = 0.5, fill = "#b2abd2") +
annotate("rect", xmin = 0.5, ymin = 2.5, xmax = 1, ymax = Inf, alpha = 0.5, fill = "#5e3c99") +
geom_text() +
geom_point() +
geom_label(data = labels, colour = "white", fill = "black") +
xlim(0, 1)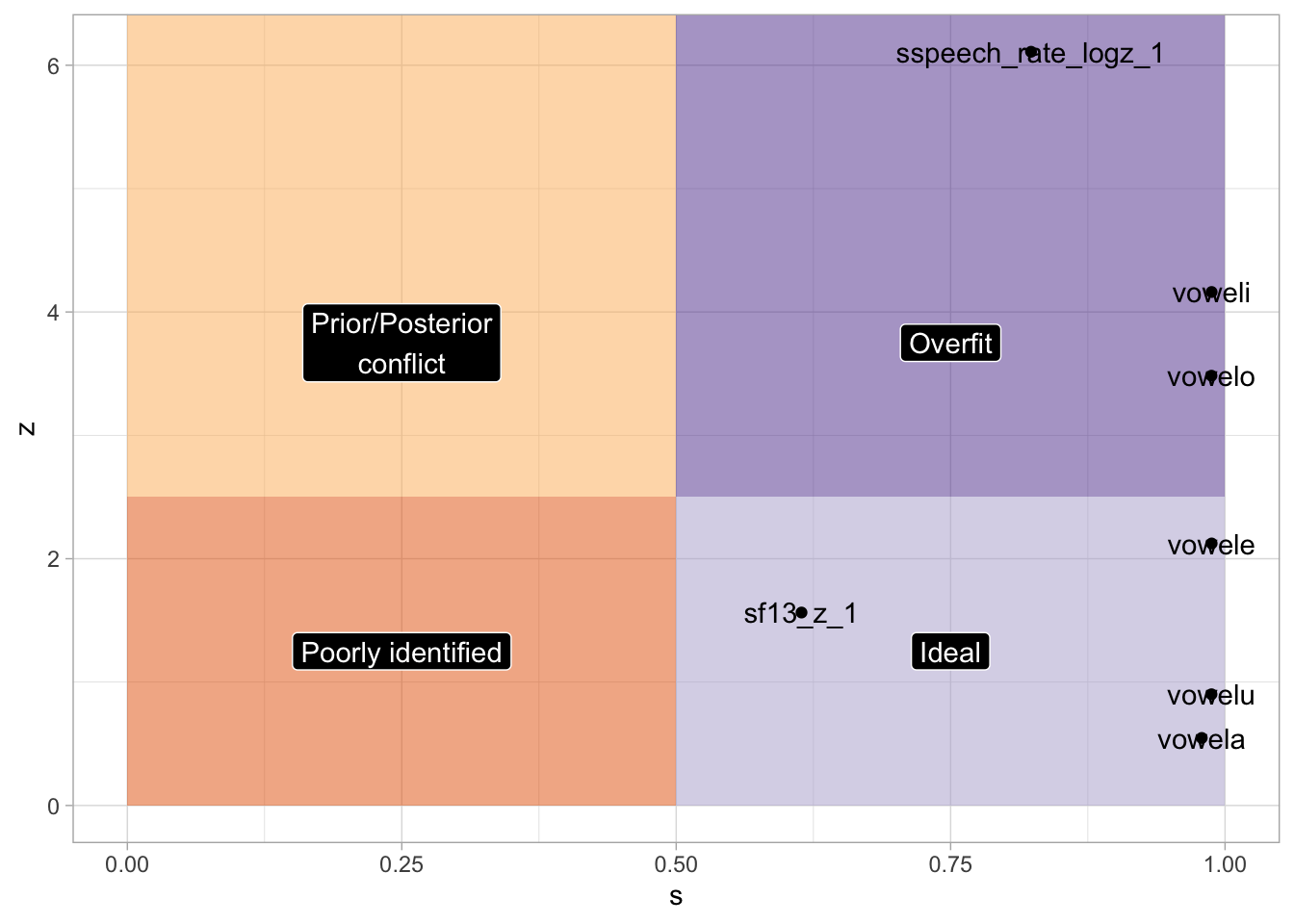
6.4 Model plotting
plot(conditional_effects(bms_1, "f13_z:vowel"), plot = FALSE)[[1]] + facet_wrap(~vowel)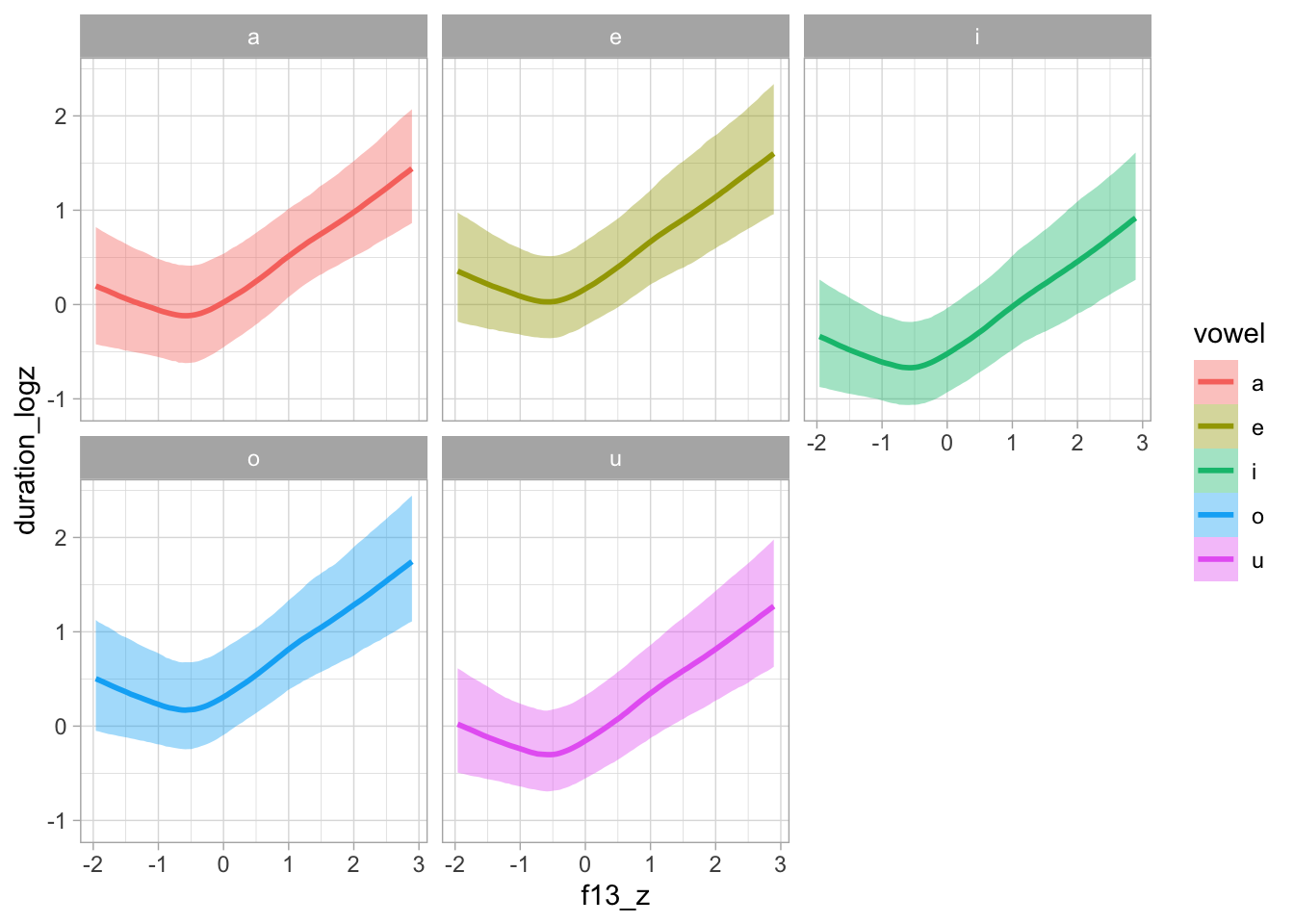
plot(conditional_effects(bms_1, "f13_z:vowel", spaghetti = TRUE, ndraws = 100), plot = FALSE)[[1]] + facet_wrap(~vowel)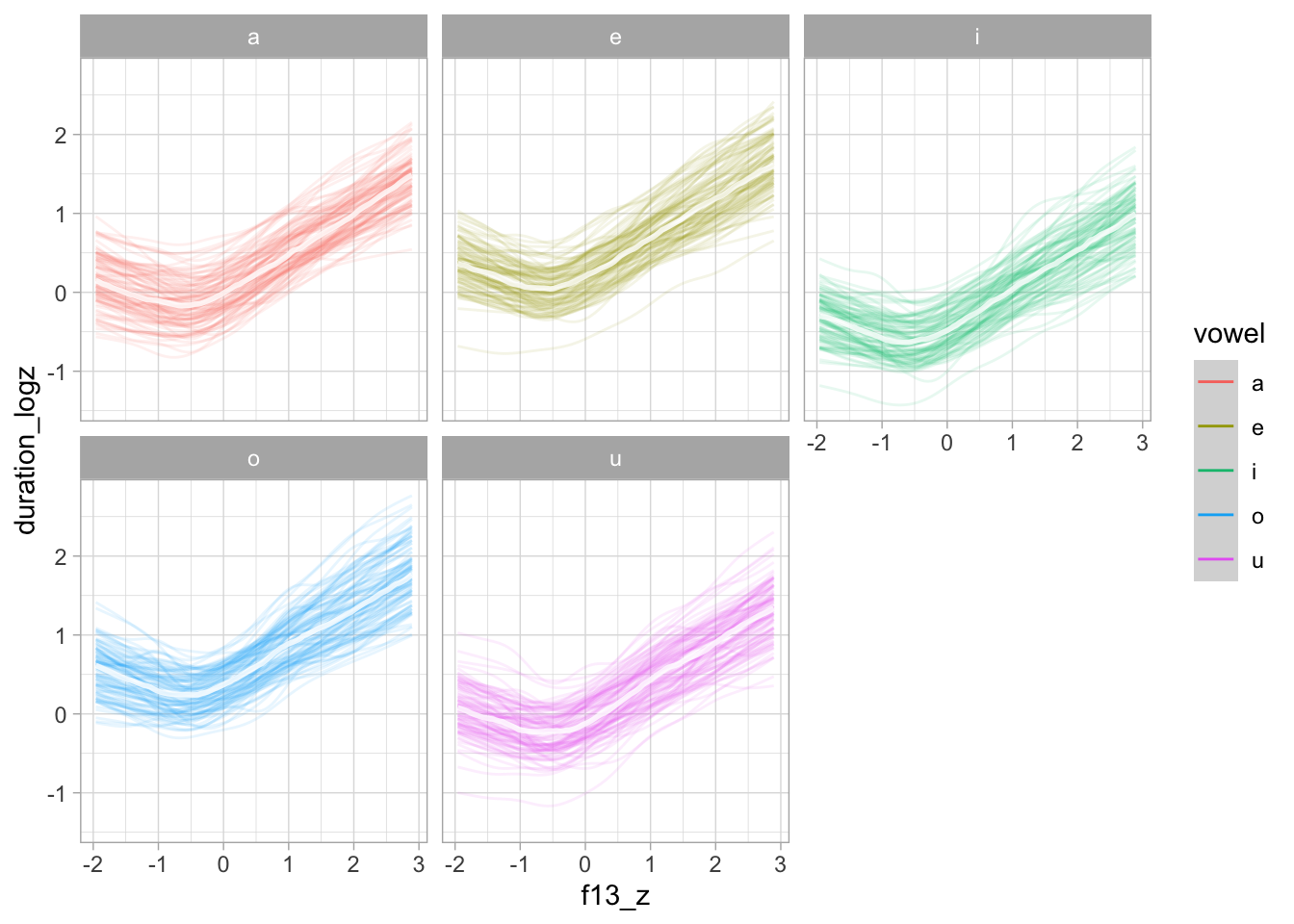
Let’s plot on the original scale.
seq_minmax <- function(x, by = 1) {
seq(min(x), max(x), by = by)
}
bms_1_grid <- expand_grid(
vowel = levels(ita_egg$vowel),
f13_z = seq_minmax(ita_egg$f13_z, 0.25),
speech_rate_logz = 0,
speaker = NA
)
bms_1_preds <- epred_draws(bms_1, newdata = bms_1_grid, re_formula = NA) |>
mutate(
duration_log = .epred * sd(ita_egg$duration_log) + mean(ita_egg$duration_log),
duration = exp(duration_log),
f13 = f13_z * sd(ita_egg$f13) + mean(ita_egg$f13)
)bms_1_grid_m <- expand_grid(
vowel = levels(ita_egg$vowel),
f13_z = 0,
speech_rate_logz = 0,
speaker = NA
)
bms_1_preds_m <- epred_draws(bms_1, newdata = bms_1_grid_m, re_formula = NA) |>
mutate(
duration_log = .epred * sd(ita_egg$duration_log) + mean(ita_egg$duration_log),
duration = exp(duration_log),
f13 = f13_z * sd(ita_egg$f13) + mean(ita_egg$f13)
)
mean_pred_vdur <- round(mean(bms_1_preds_m$duration))
bms_1_preds |>
mutate(
vowel = str_replace(vowel, "o", "ɔ"),
vowel = factor(vowel, levels = c("i", "u", "e", "ɔ", "a"))
) |>
group_by(vowel, f13) |>
ggplot(aes(f13, duration, fill = vowel)) +
geom_hline(yintercept = mean_pred_vdur, linetype = "dashed", colour = "red") +
stat_ribbon(.width = 0.98, alpha = 0.2) +
stat_ribbon(.width = 0.9, alpha = 0.4) +
stat_lineribbon(.width = 0.6, alpha = 0.8) +
geom_vline(data = vmean_f13 |> mutate(
vowel = str_replace(vowel, "o", "ɔ"),
vowel = factor(vowel, levels = c("i", "u", "e", "ɔ", "a"))
), aes(xintercept = f13_mean, colour = vowel), linetype = "dashed") +
geom_rug(data = ita_egg |> mutate(
vowel = str_replace(vowel, "o", "ɔ"),
vowel = factor(vowel, levels = c("i", "u", "e", "ɔ", "a"))
), alpha = 0.1, length = unit(0.015, "npc"), aes(colour = vowel)) +
facet_wrap(~vowel) +
labs(
x = "F1 (Hz)", y = "Duration (ms)"
) +
scale_fill_manual(values = cols) +
scale_colour_manual(values = cols) +
theme(legend.position = "none")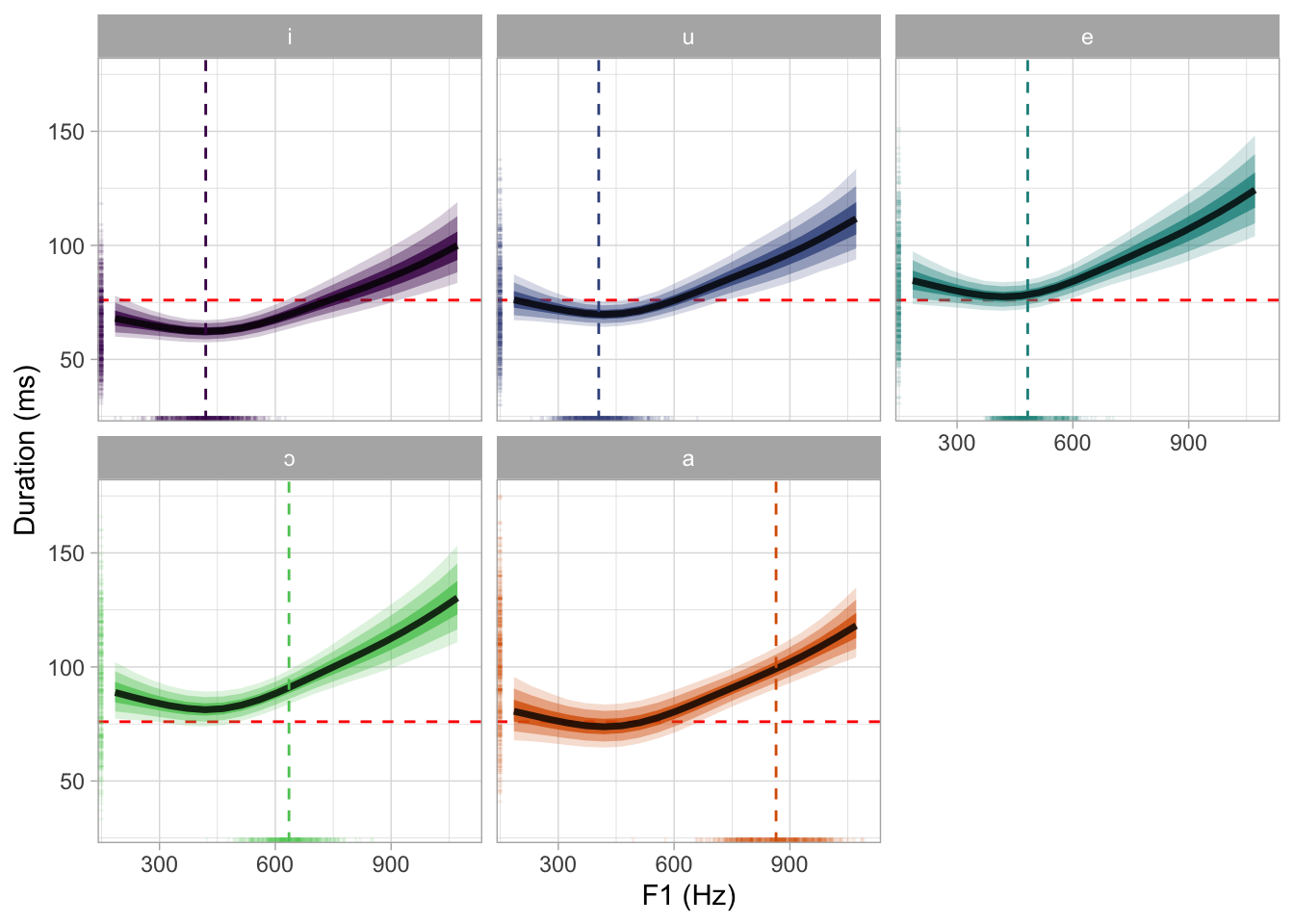
ggsave("img/bms1-pred-plot-ms-hz.png", width = 7, height = 5)7 Non-linear modelling: mgcv F1 and F2
This is extra modelling of F1/F2 using mgcv. Does not feature in the publication and it’s unrelated to the research questions of the paper.
gam_2 <- bam(
duration_logz ~
vowel +
s(f13_z, f23_z) +
s(f13_z, f23_z, speaker, bs = "fs", m = 1),
data = ita_egg
)summary(gam_2)
Family: gaussian
Link function: identity
Formula:
duration_logz ~ vowel + s(f13_z, f23_z) + s(f13_z, f23_z, speaker,
bs = "fs", m = 1)
Parametric coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.03176 0.18200 -0.175 0.861477
vowele 0.43854 0.15991 2.742 0.006137 **
voweli -0.31252 0.16553 -1.888 0.059128 .
vowelo 0.38534 0.10901 3.535 0.000414 ***
vowelu -0.11135 0.15542 -0.716 0.473775
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Approximate significance of smooth terms:
edf Ref.df F p-value
s(f13_z,f23_z) 12.89 16.78 4.814 <2e-16 ***
s(f13_z,f23_z,speaker) 102.60 567.00 6.872 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
R-sq.(adj) = 0.708 Deviance explained = 71.9%
fREML = 2575.1 Scale est. = 0.29223 n = 3019gam_2_preds <- predict_gam(gam_2, length_out = 50, exclude_terms = "s(f13_z,f23_z,speaker)")vmeans <- ita_egg |>
group_by(vowel) |>
summarise(
f13_z = mean(f13_z), f23_z = mean(f23_z)
)
gam_2_preds |>
ggplot(aes(f23_z, f13_z)) +
geom_raster(aes(fill = duration_logz), interpolate = TRUE) +
geom_contour(aes(z = duration_logz), bins = 40, colour = "white", linewidth = 0.05) +
geom_label(data = vmeans, aes(label = vowel), size = 5) +
scale_x_reverse(position = "top") +
scale_y_reverse(position = "right") +
scale_fill_distiller(palette = "BuPu")Warning: Contour data has duplicated x, y coordinates.
ℹ 244494 duplicated rows have been dropped.